Life is about Connections
Friends of the Foundary & Biomaths Colloquium
John "Scooter" Morris, Ph.D.
Swansea University
19 June 2019
Slides: https://rbvi.github.io/chimera-tutorials/presentations/FriendsOfTheFoundary-2019.html/
Connections
How did I get here?

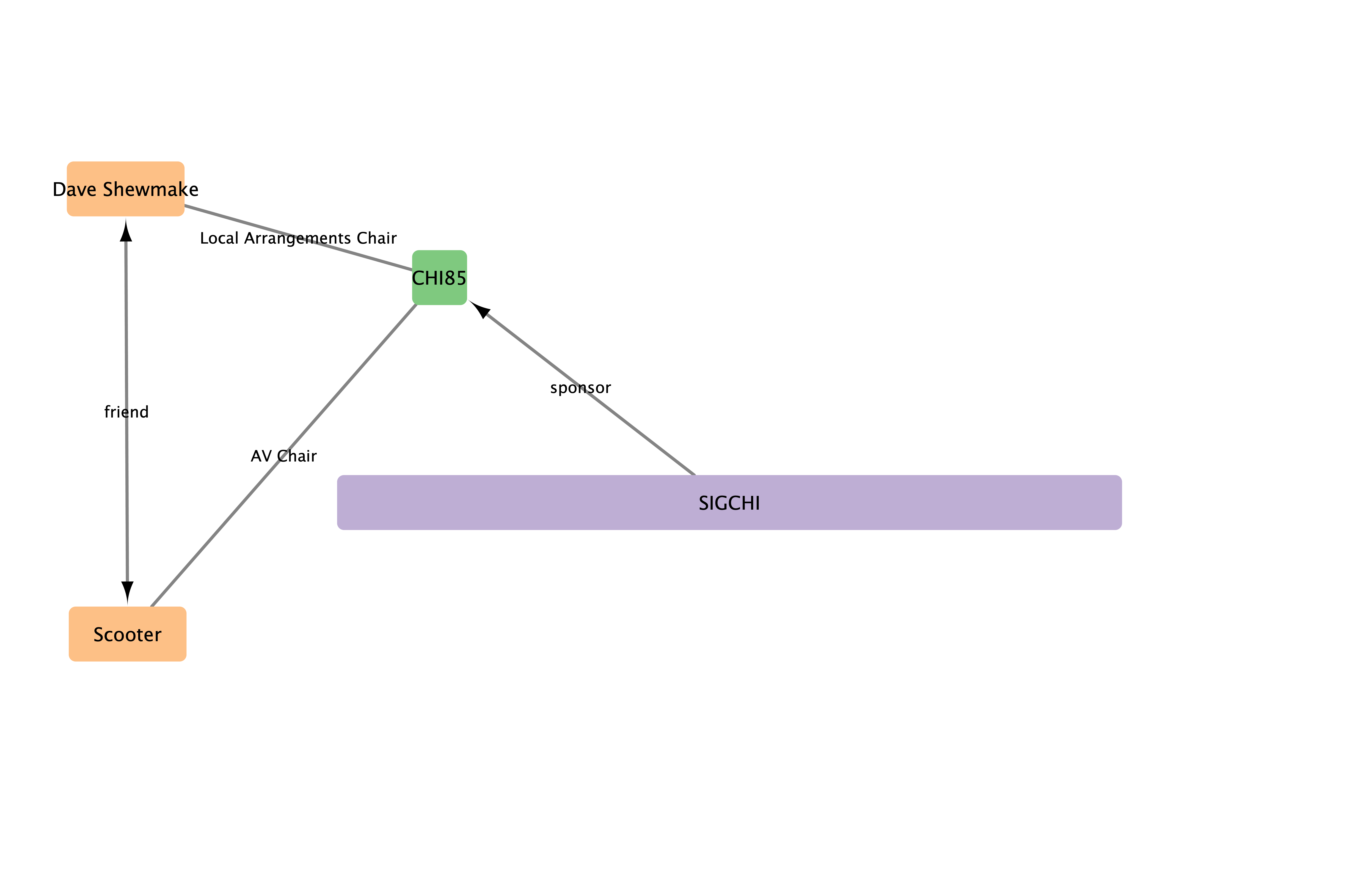
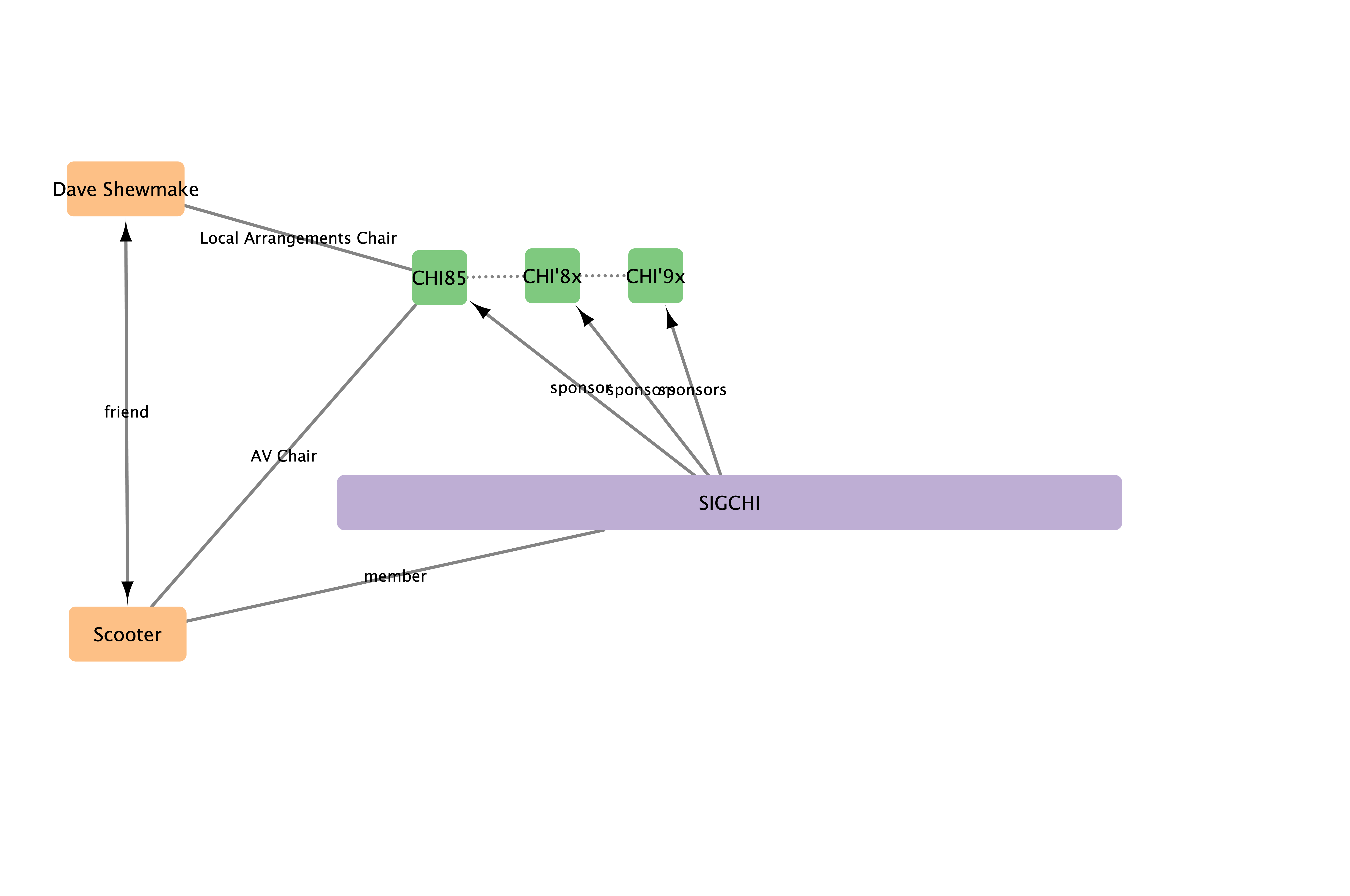
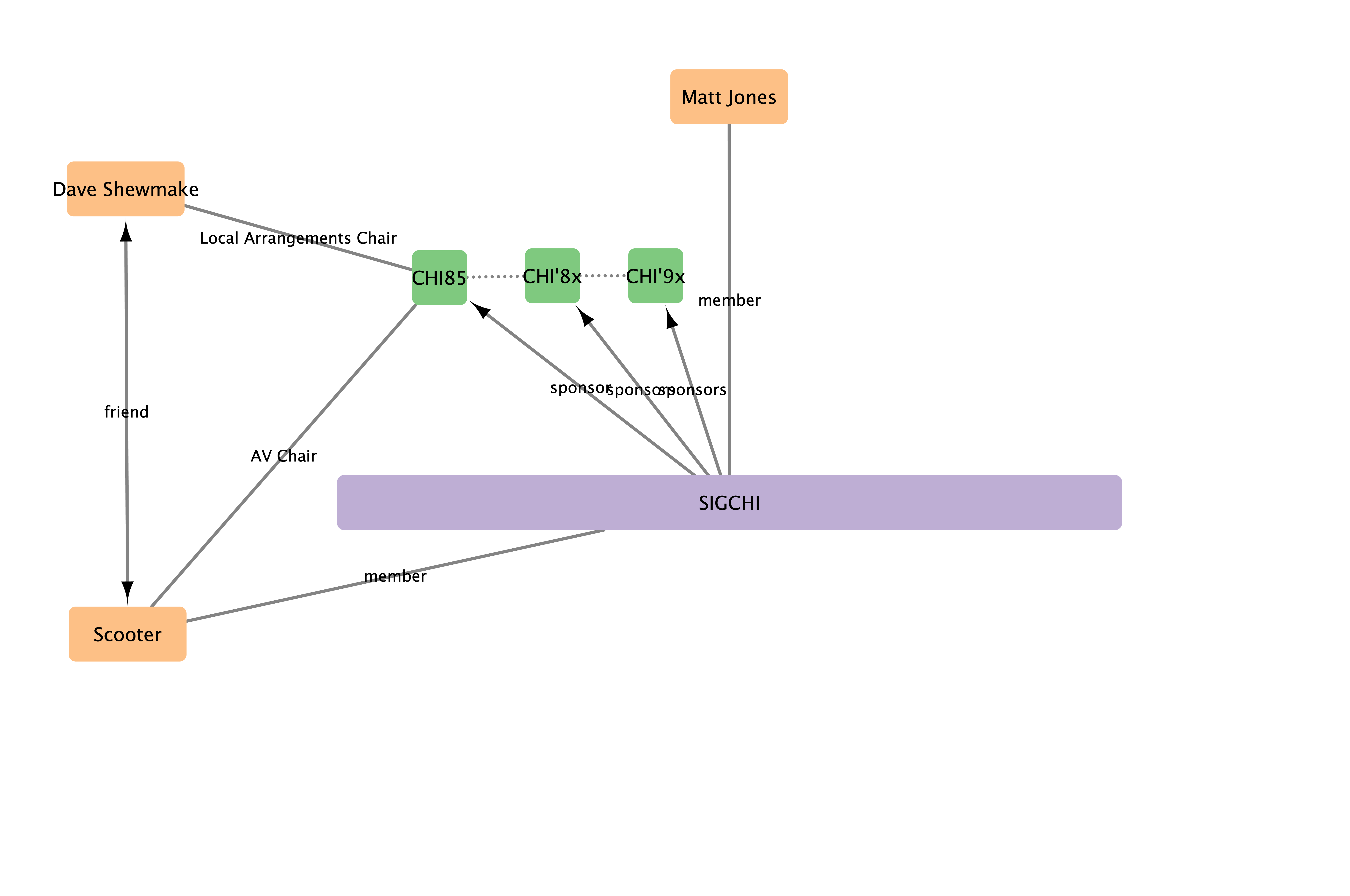
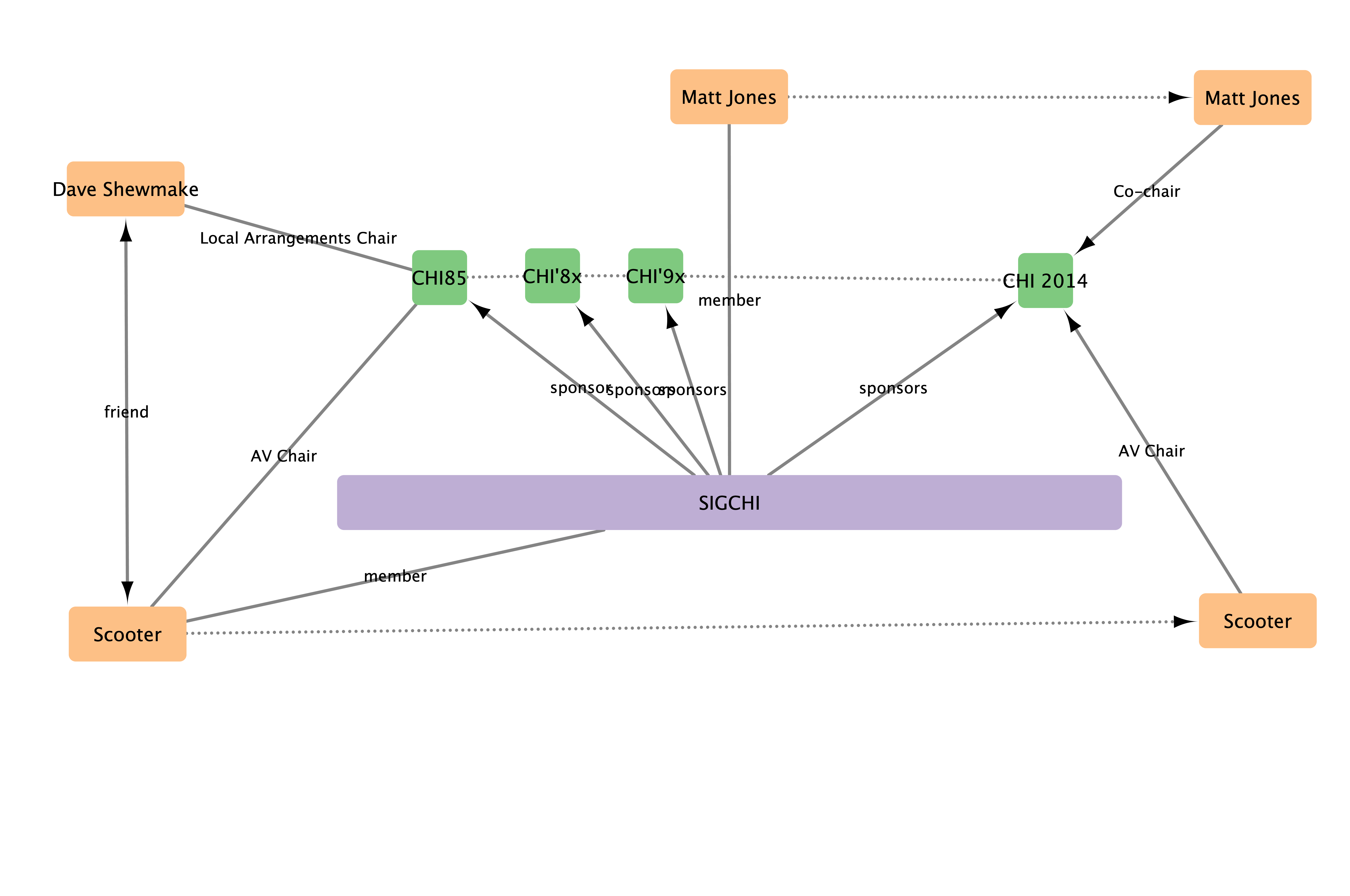
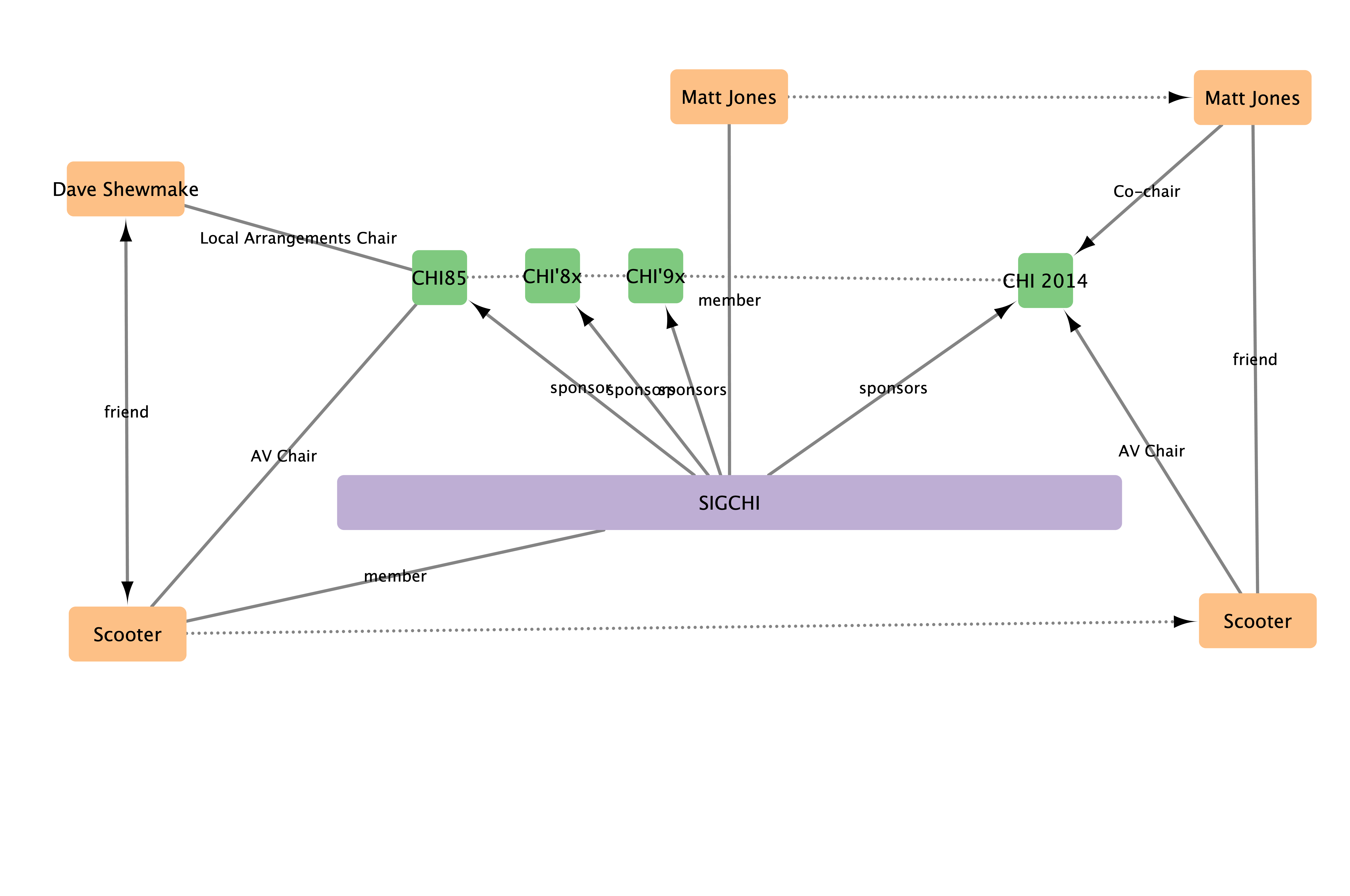
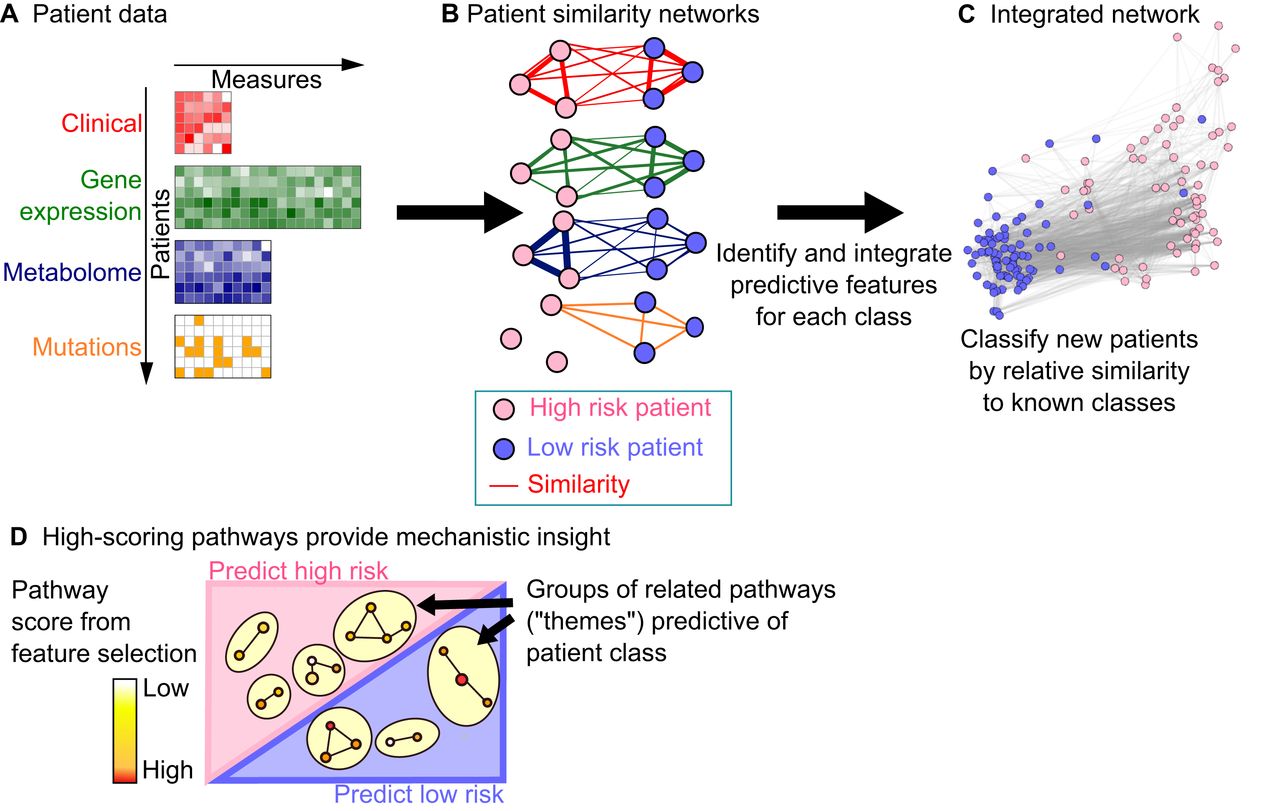
Connections
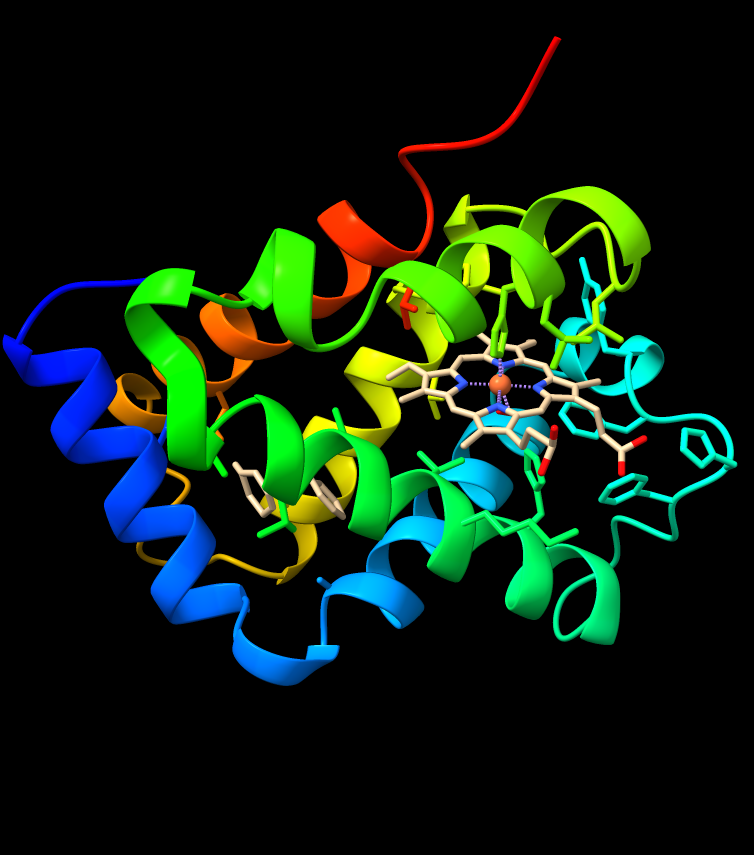
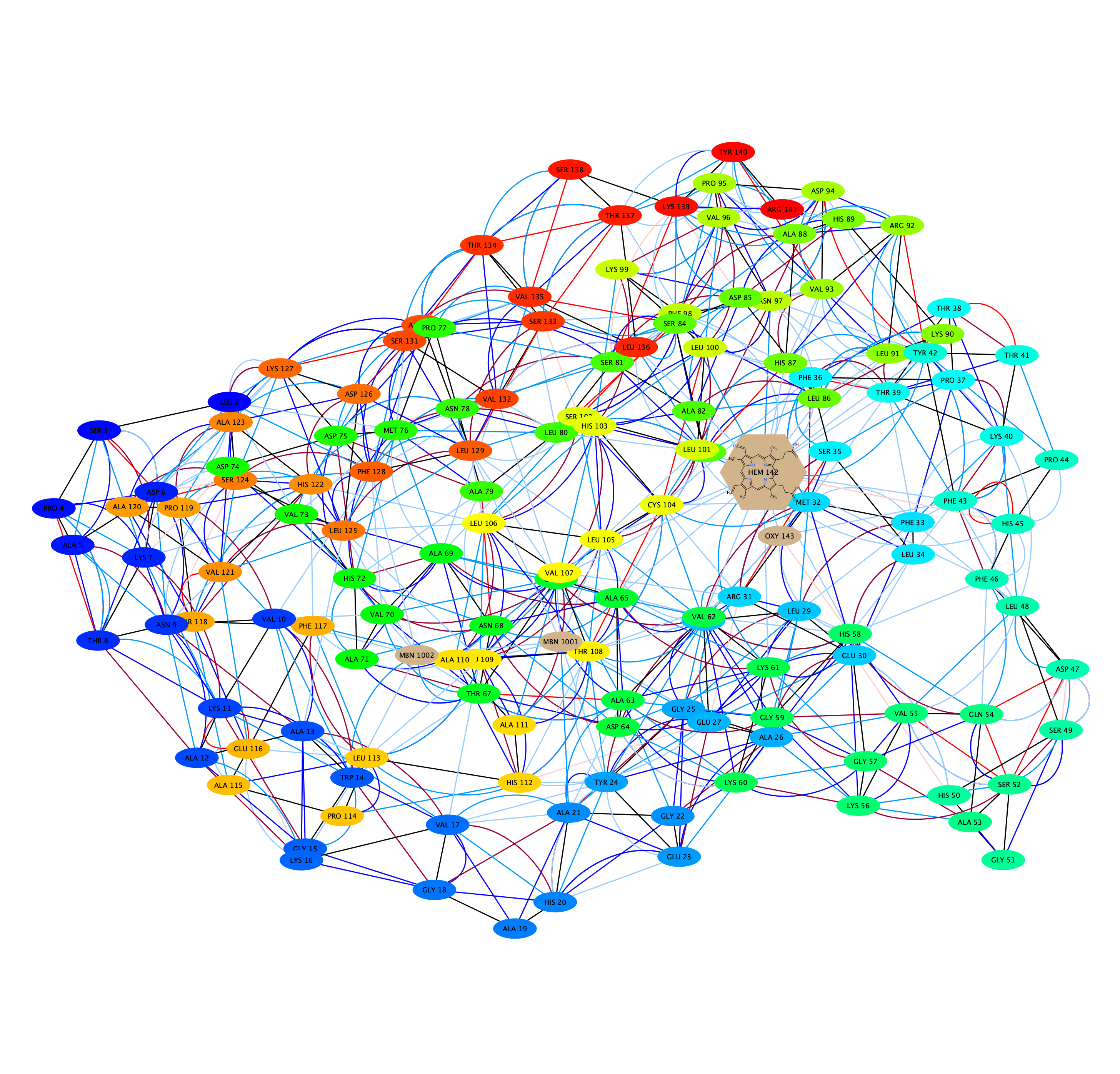
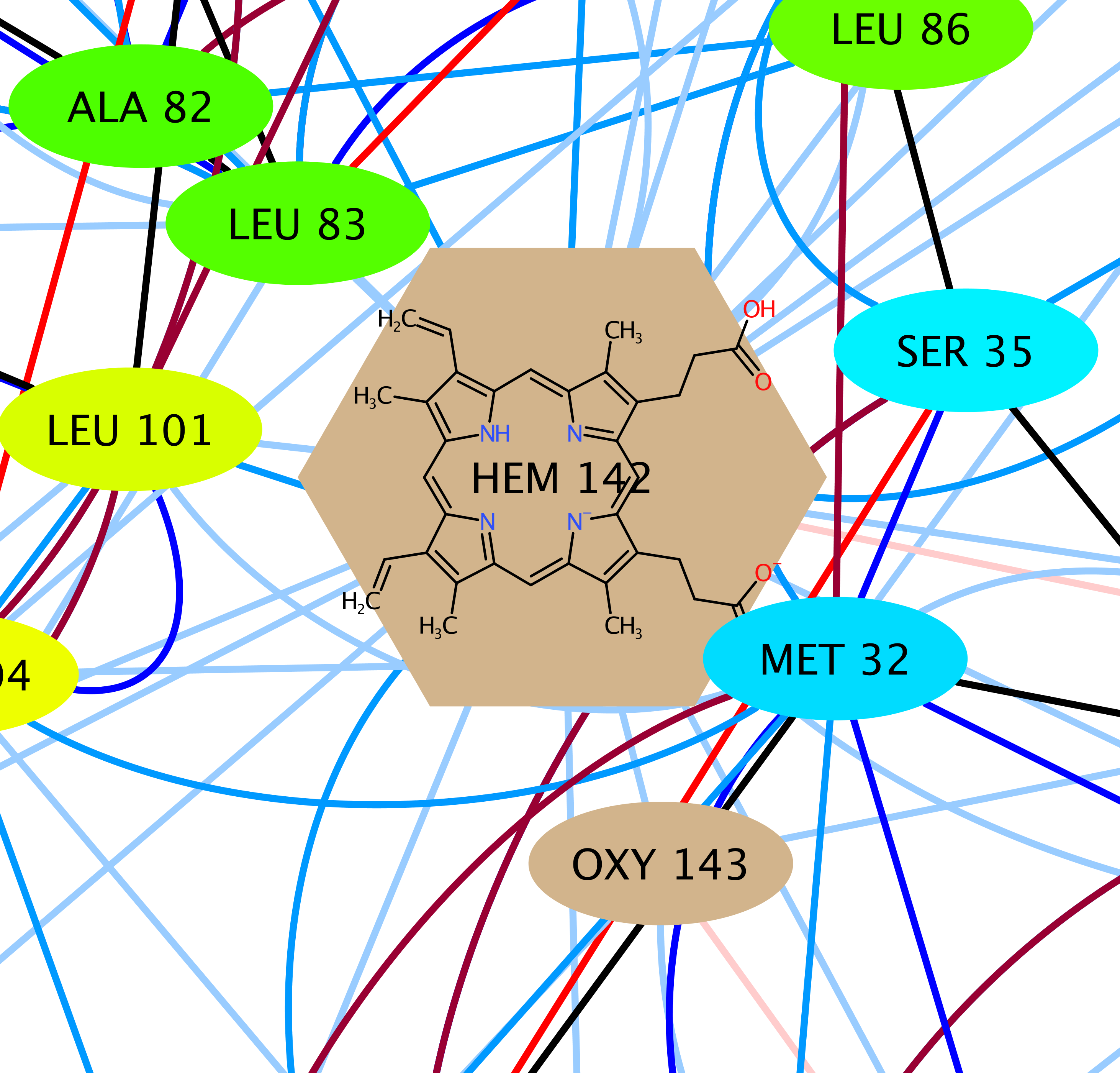
Connections
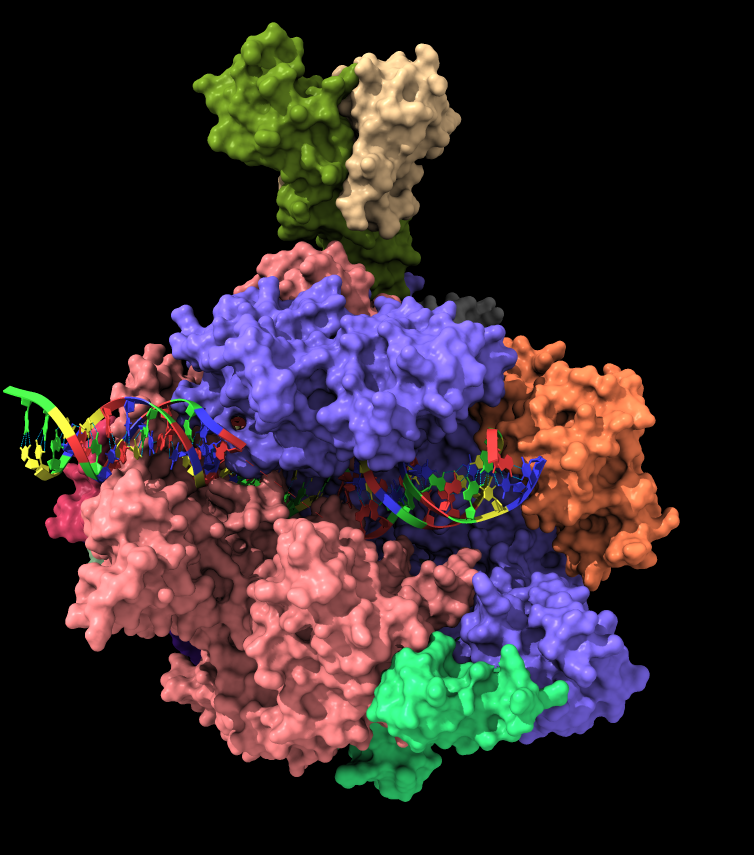
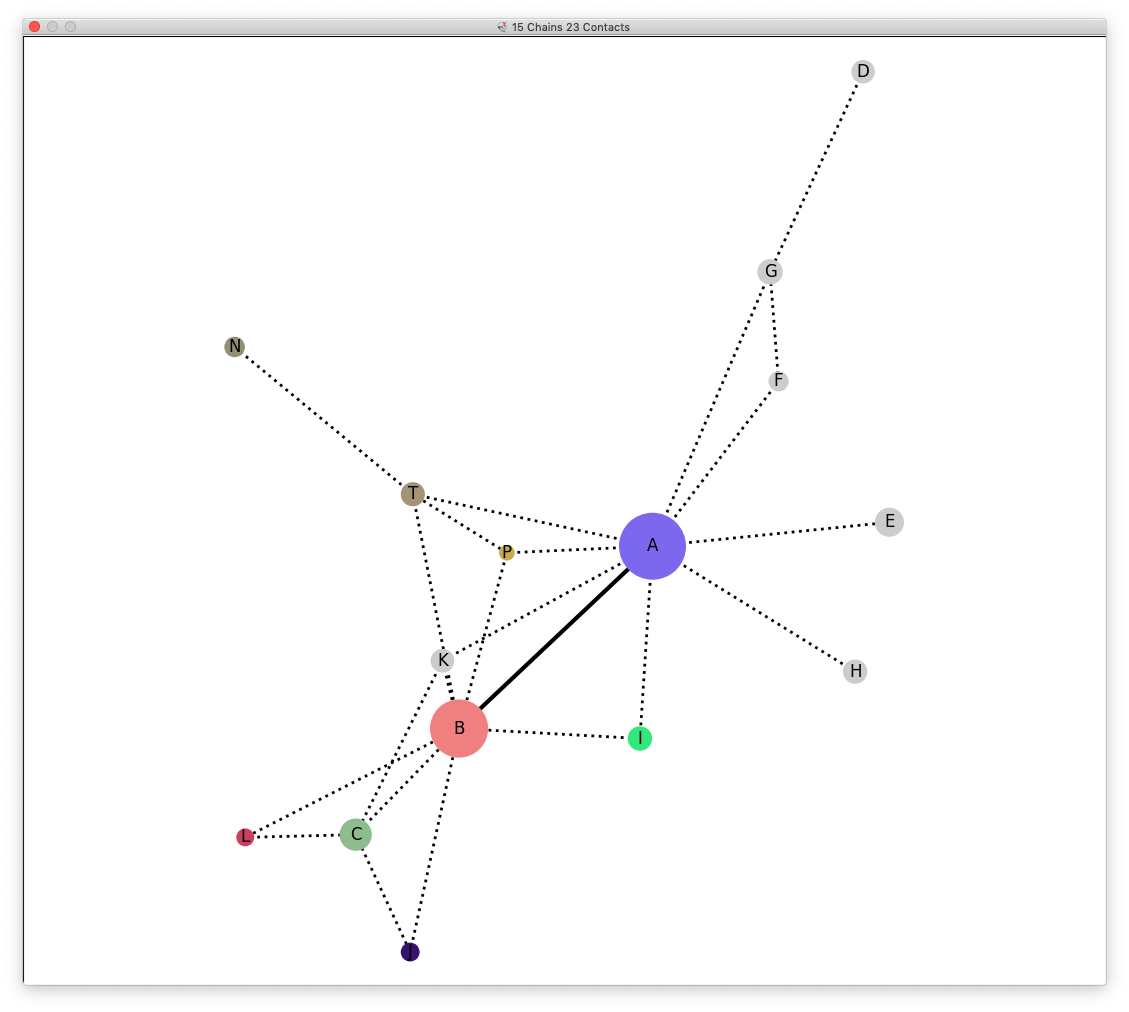
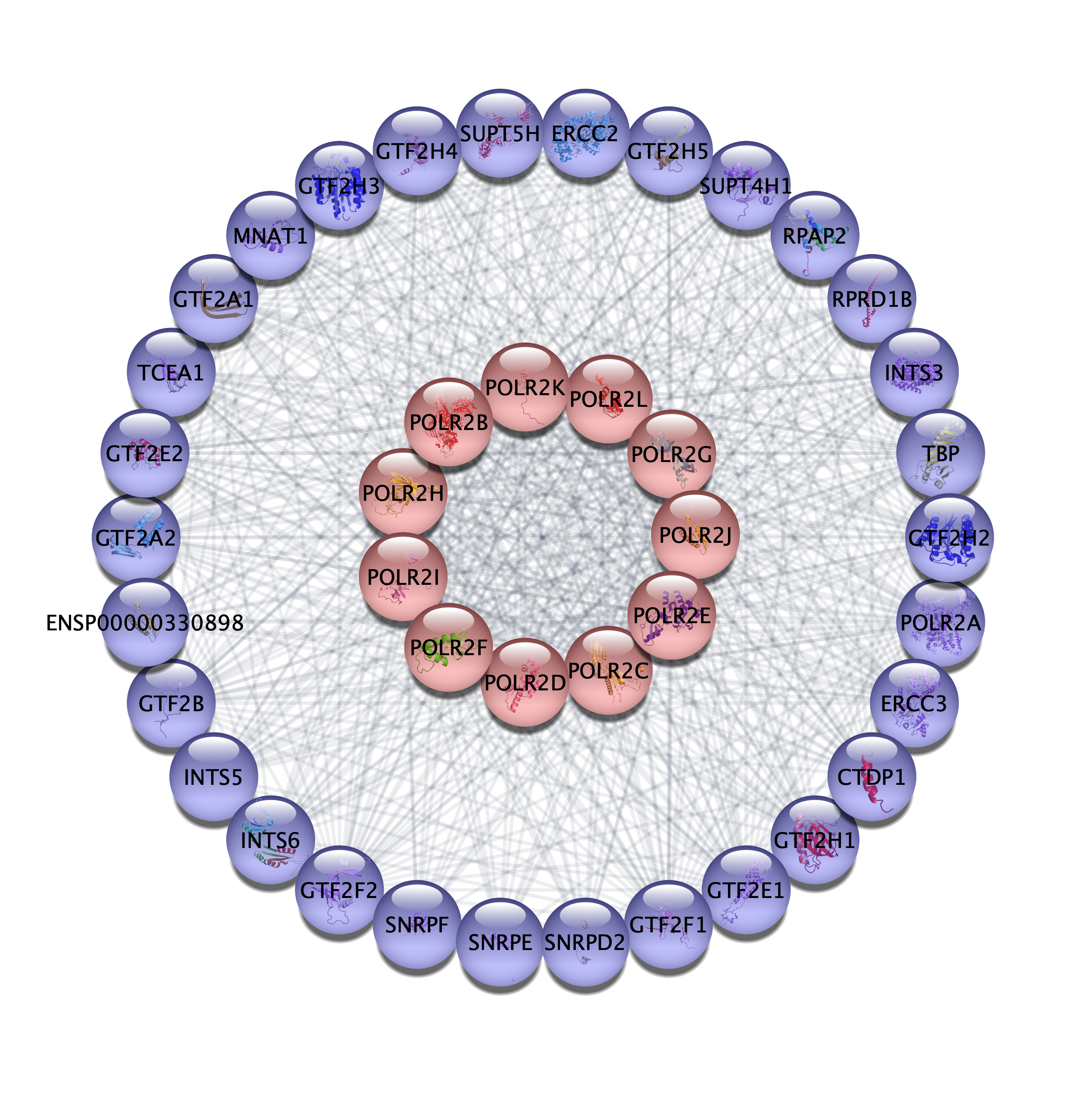
Connections
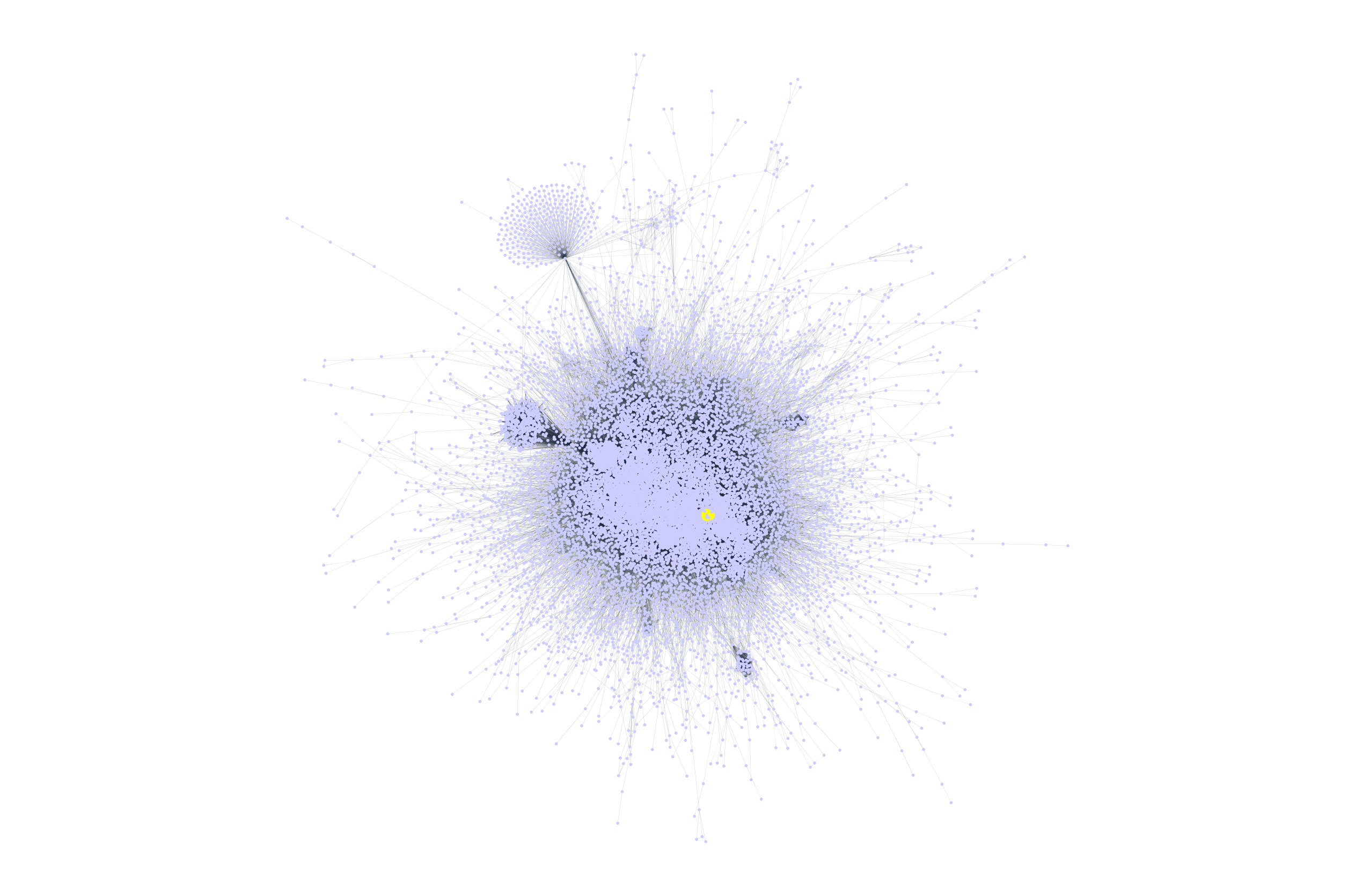
Connections

Connections
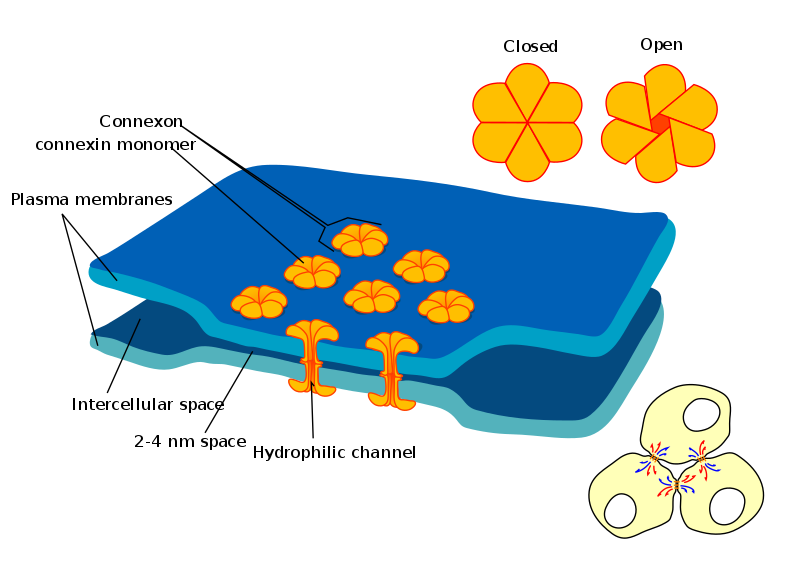
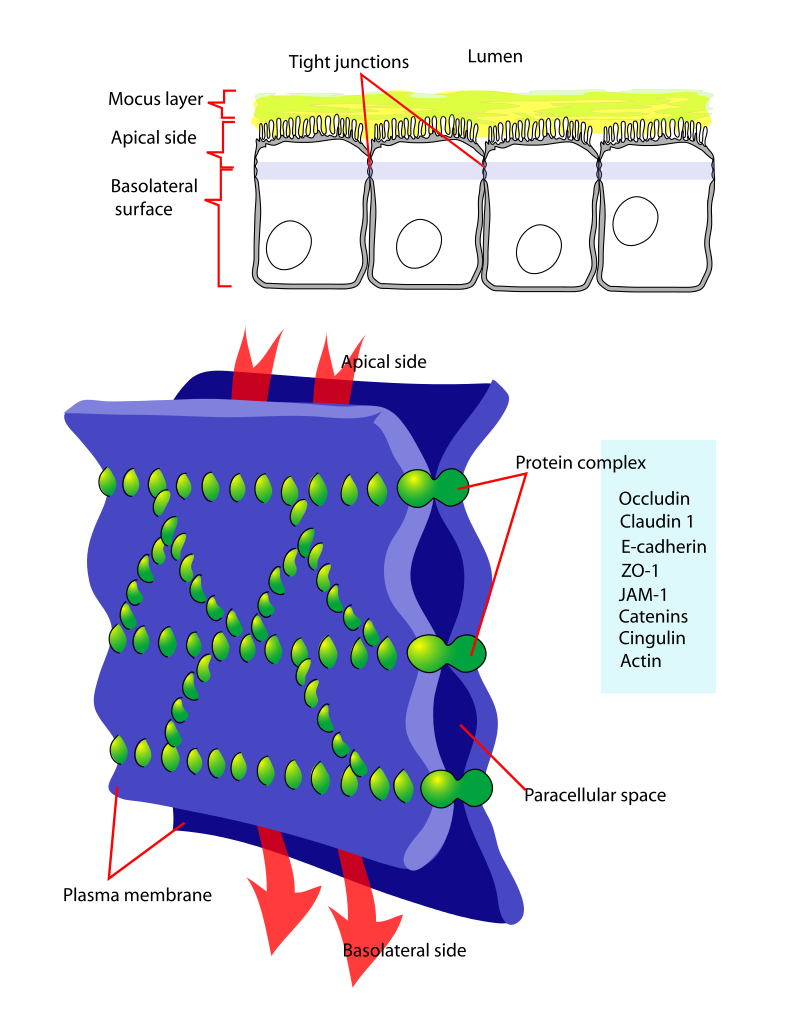
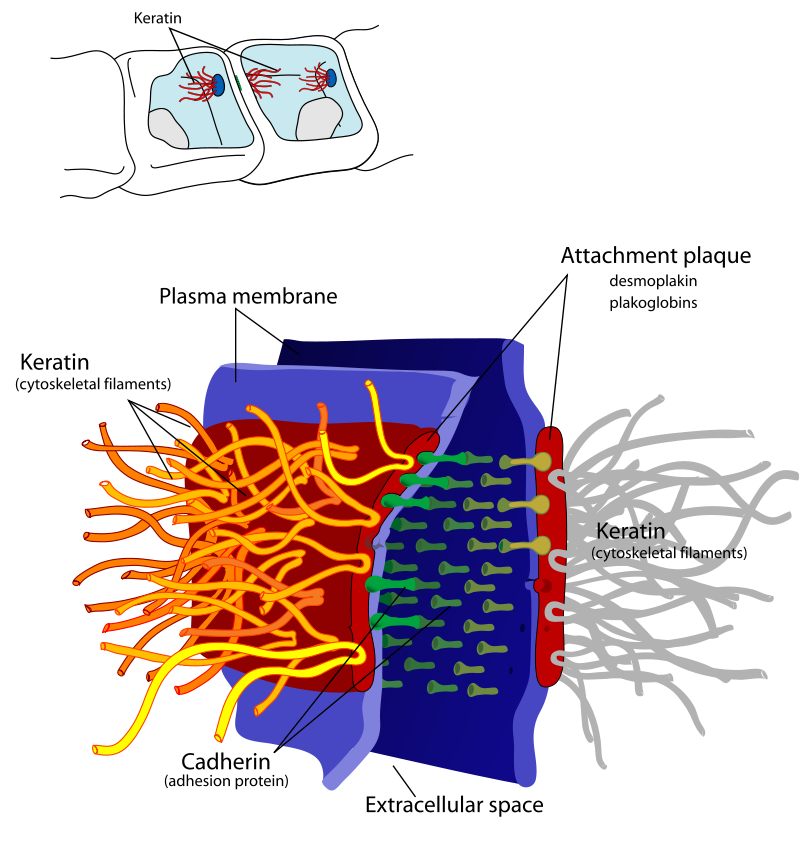
- Cell-cell interactions
- Gap junctions
- Tight junctions
- Desmosomes
- Cell signaling
Connections - why do we care?
“...empirically derived networks are necessary for describing the molecular mechanisms and biological processes that drive disease under the influences of inherited risk factors (genetic markers) and environmental risk factors.”
Schadt EE, Björkegren JL. Science translational medicine. 2012
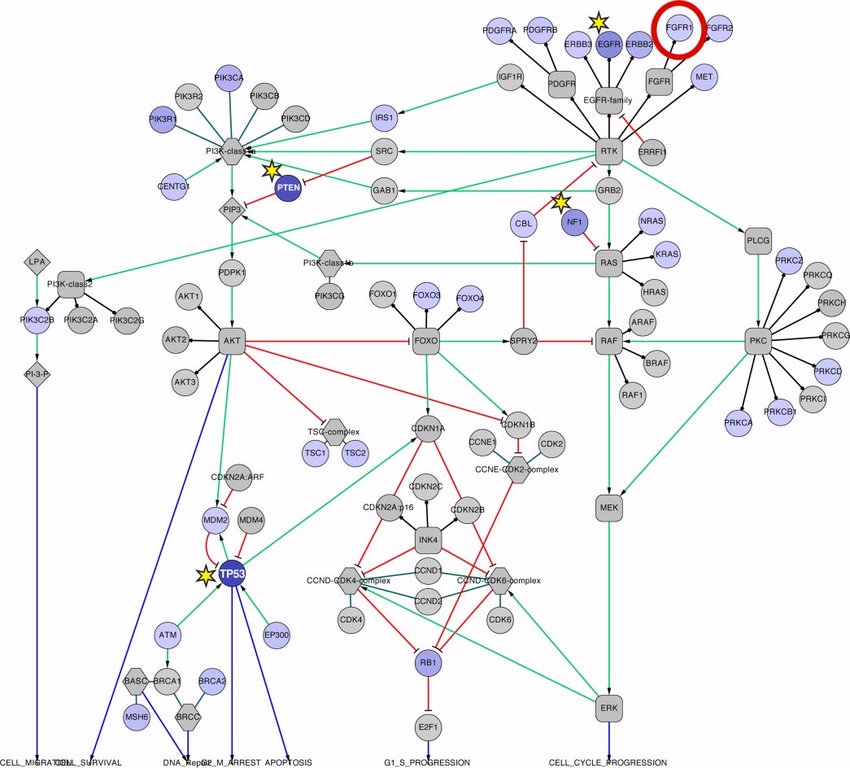
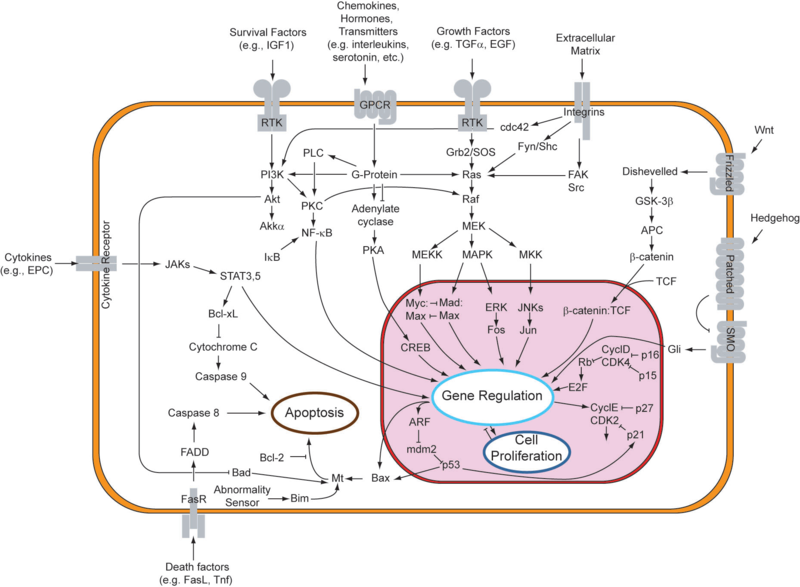
Connections - why do we care?
Finding connections
- Molecular interactions
- Structural methods (X-Ray, NMR, CryoEM, Cross-linking, docking)
- Systems approaches (binding assays, yeast two hybrid, AP/MS, BioID, Single-cell genomics)
- Cell-cell interactions
- Imaging techniques
- Microfluidic approaches
- Biochemical measurements
- Interspecies interactions
- Host-pathogen proteomics (BioID, AP/MS)
(Really) Big Data
Back of the envelope calculations for # of interactions in a human
| # of proteins in a human cell | ~1x1010 |
| # of lipids in a human cell | ~1x1010 |
| # of metabolites in a human cell | ~1x108 |
| # of RNA molecules in a human cell | ~1x108 |
| # of miRNA molecules in a human cell | ~1x104 |
| # of molecular entities* = | ~1x1040 |
| # of cells in a human | ~3.7x1013 |
| Potential interactions** = | <<1.3x10107 |
* Doesn't include carbohydrates, other small RNAs
** Actual interactions are much, much less due to cellular compartments, tissue specificity, etc., etc.
(Really) Big Data
But wait, there's more!
- Connections aren't static. They vary based on:
- time (development, aging, time of day, physiological changes)
- environmental conditions
- disease (e.g. infection, somatic mutation)
- Organisms connect also
- Host/pathogen interactions
- Symbiosis
- Competition
- Predation
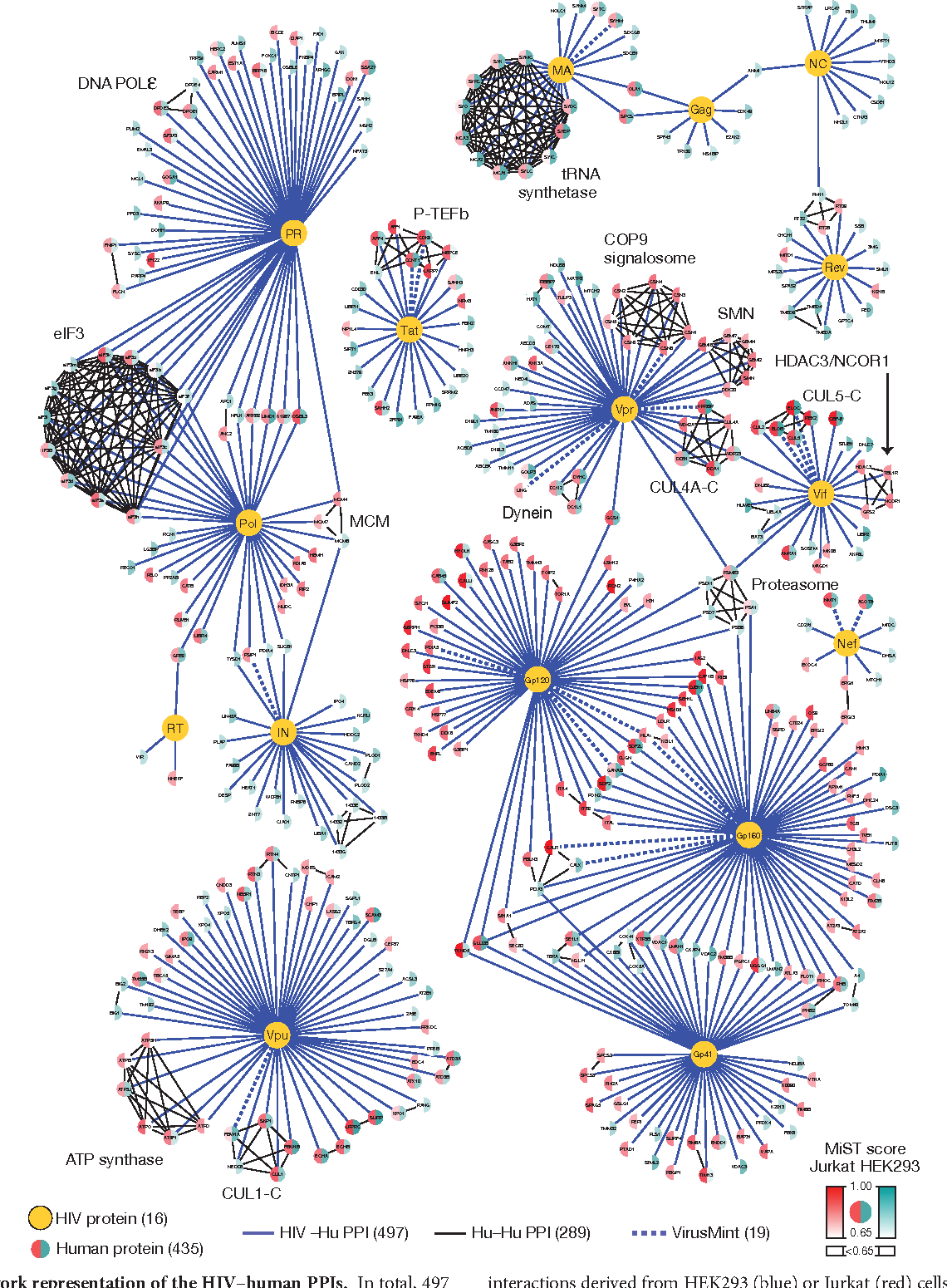
Dealing with complexity
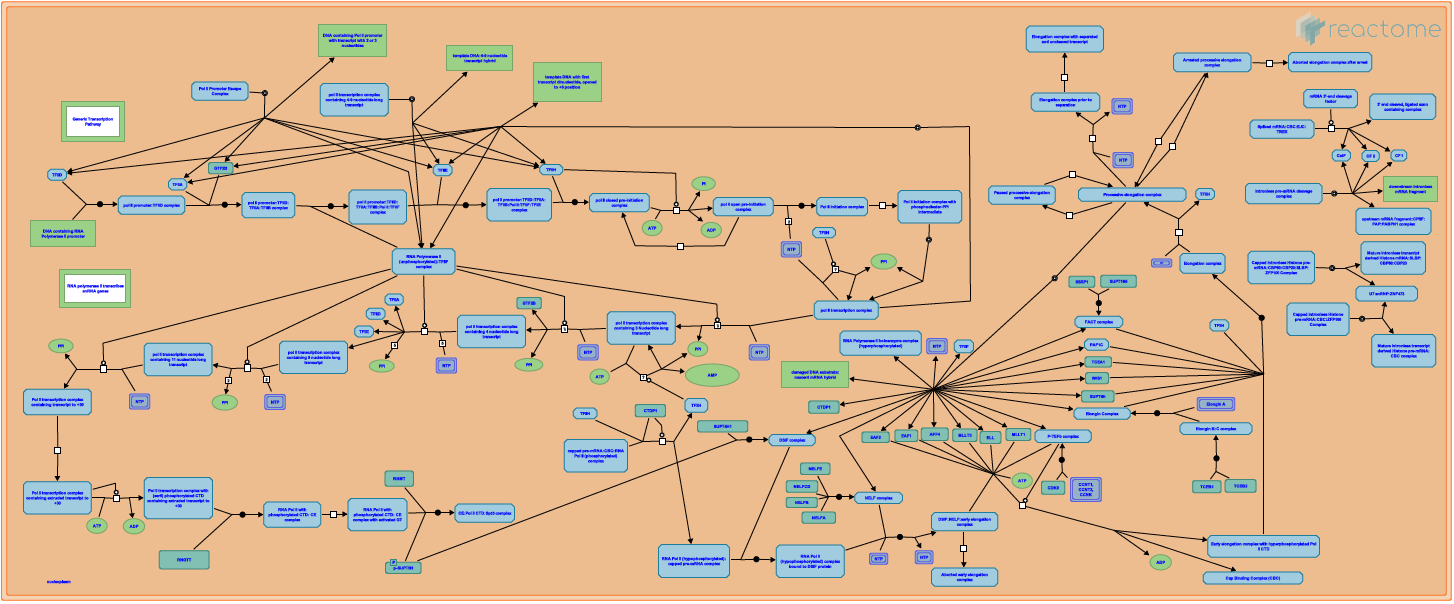
Dealing with complexity
- Topology-driven modules
- Attribute-driven modules
- Hybrid clusters
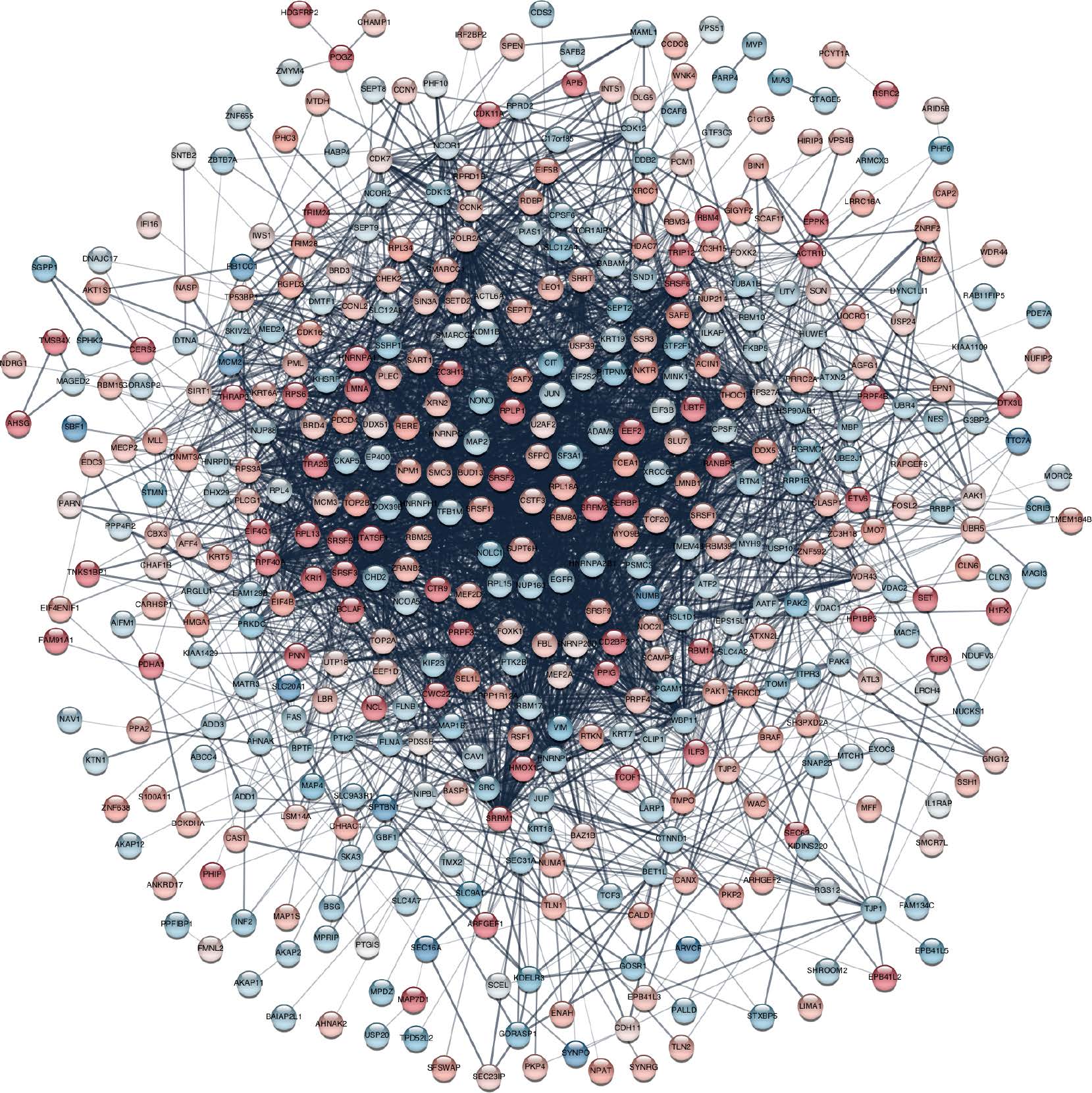
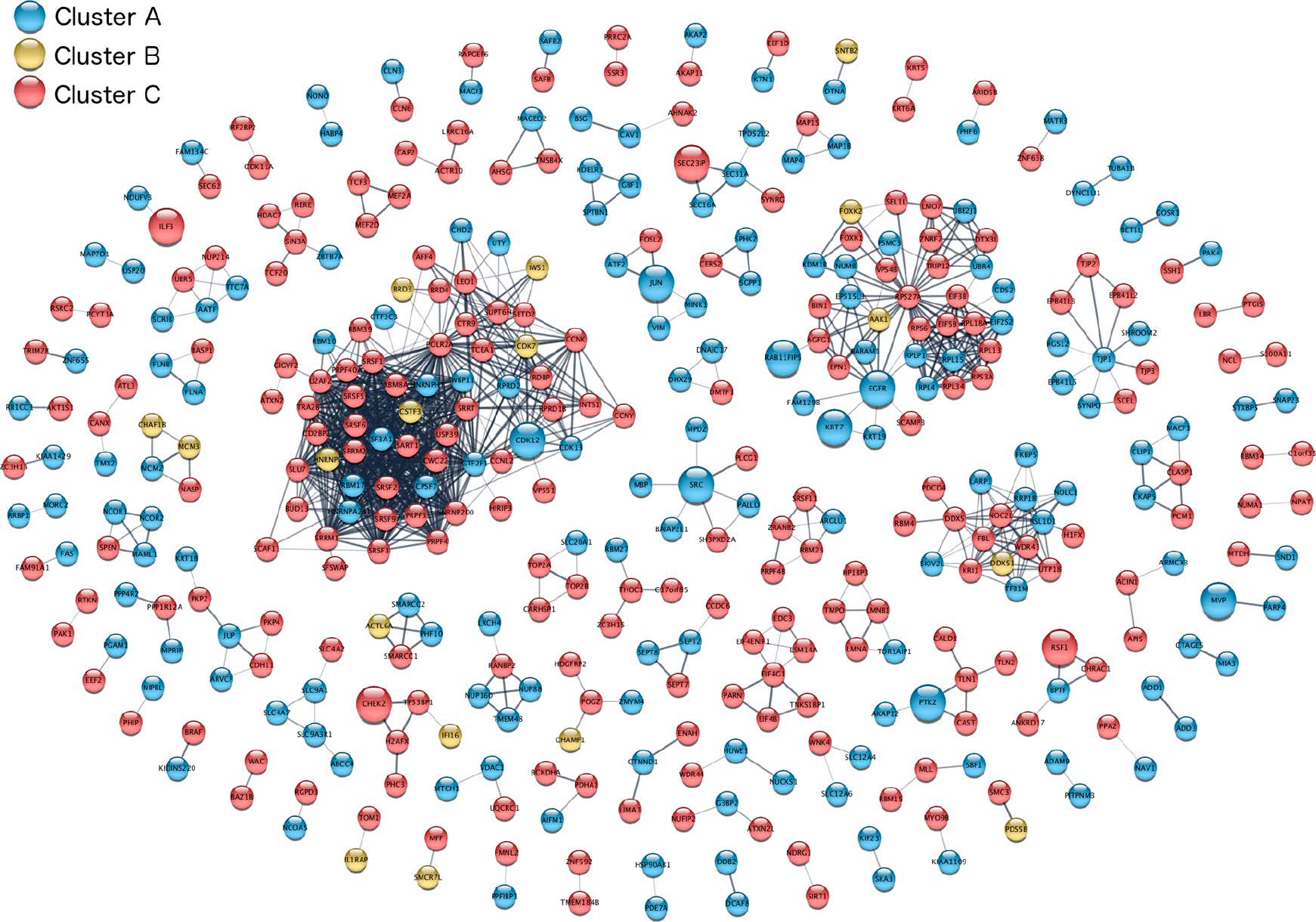
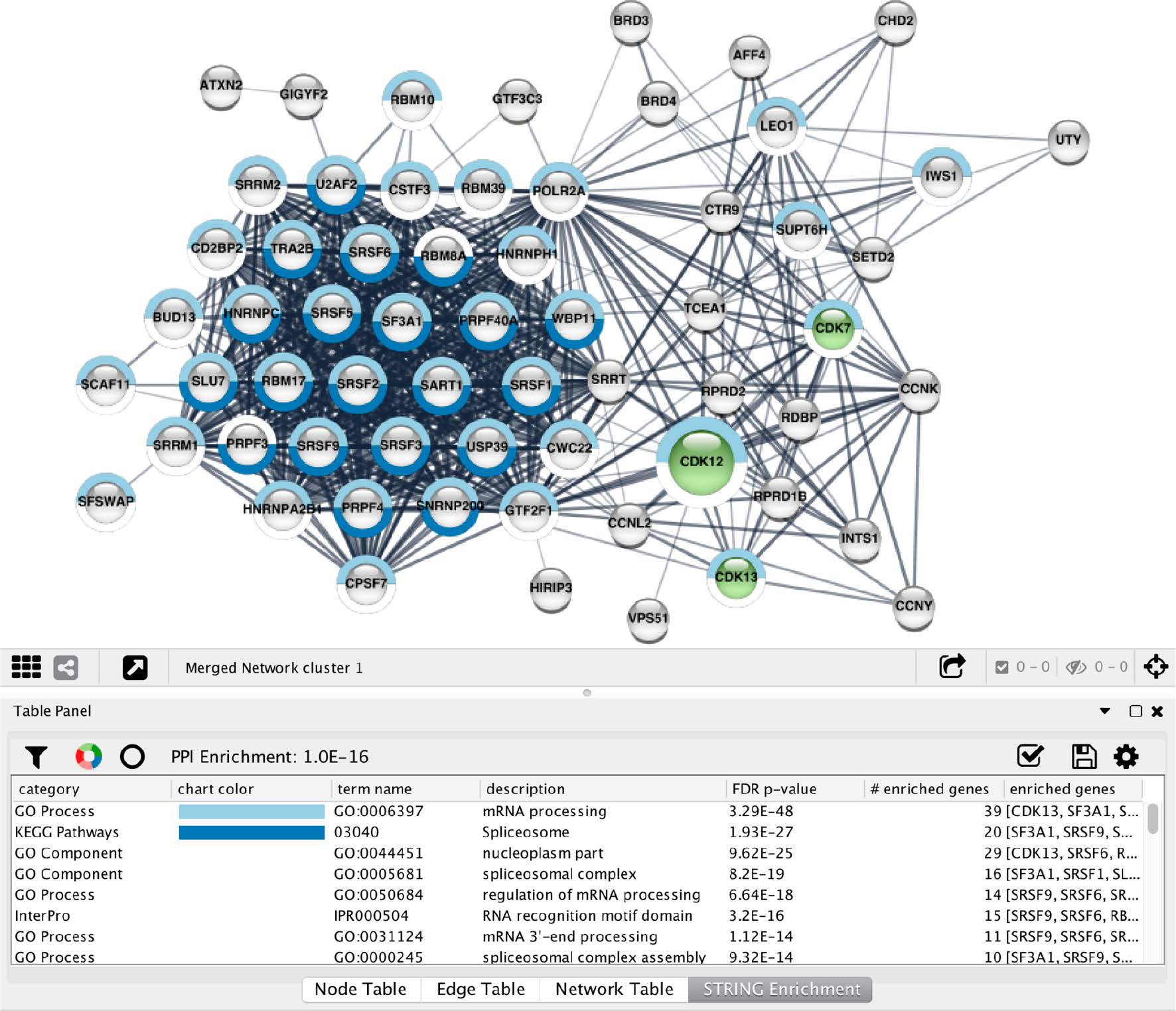
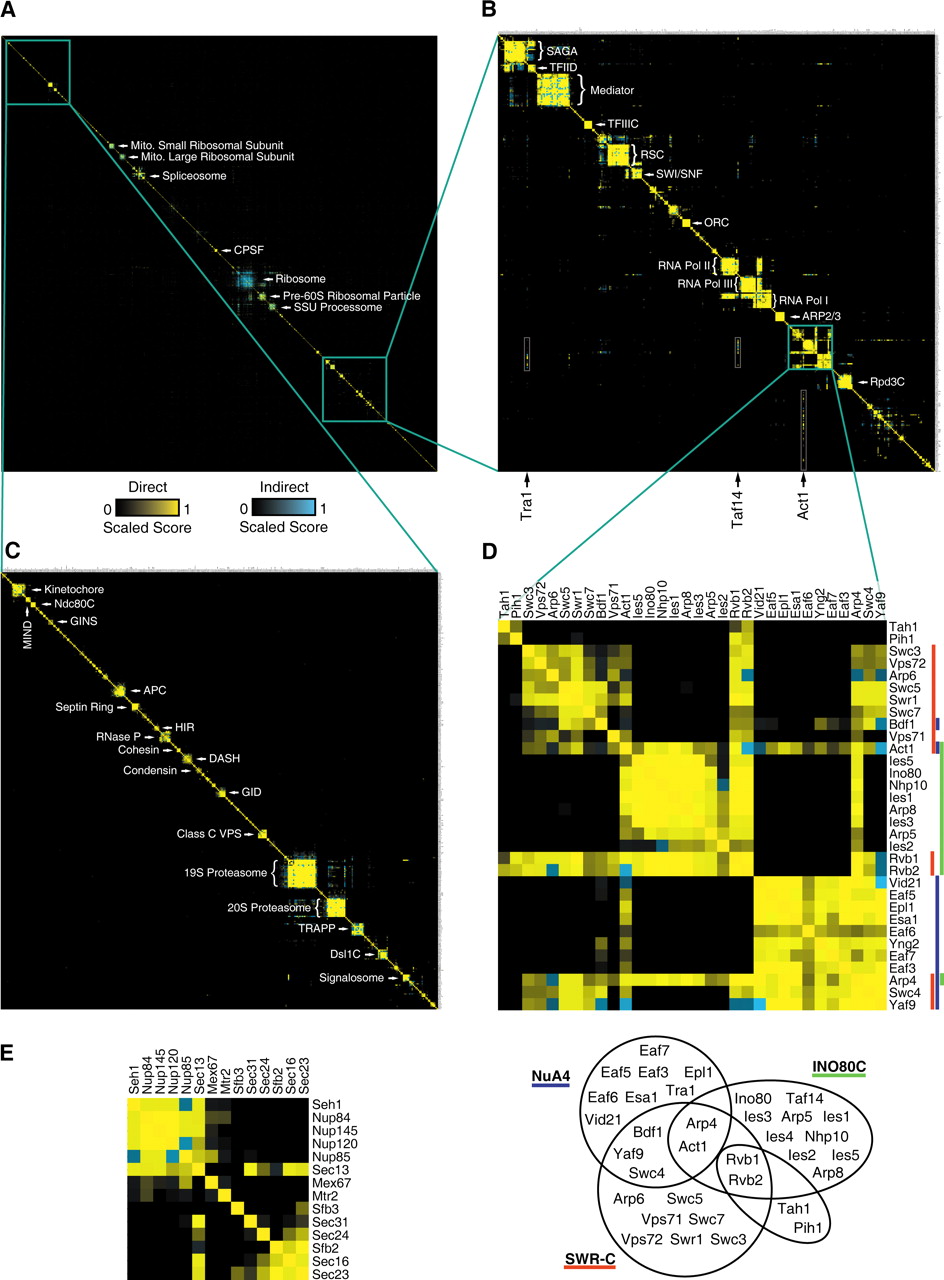

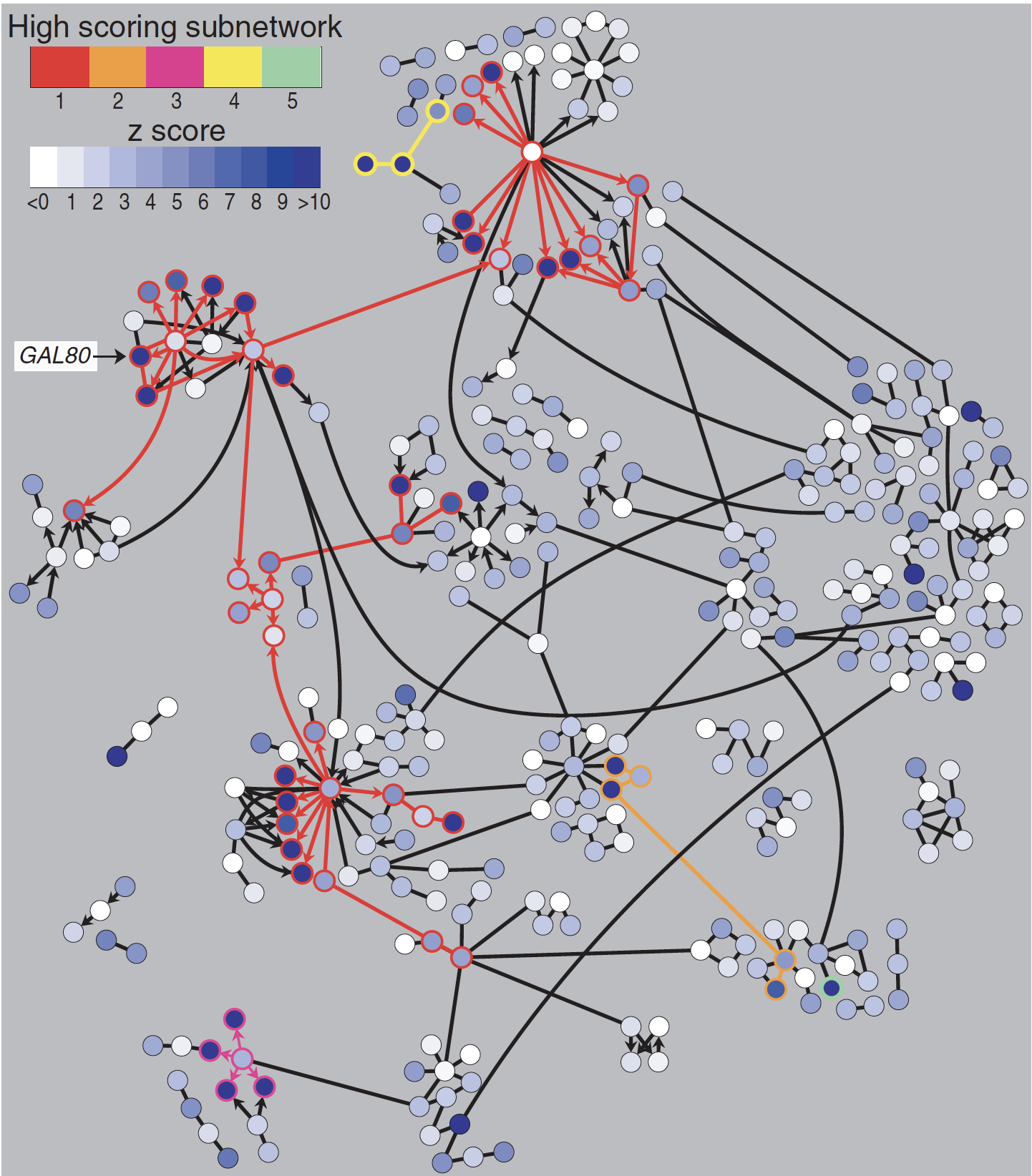
Dealing with complexity
Dealing with complexity
- Overview+detail
- Focus+context
- Linked views
- Brushing and linking
Our Projects
Tool:
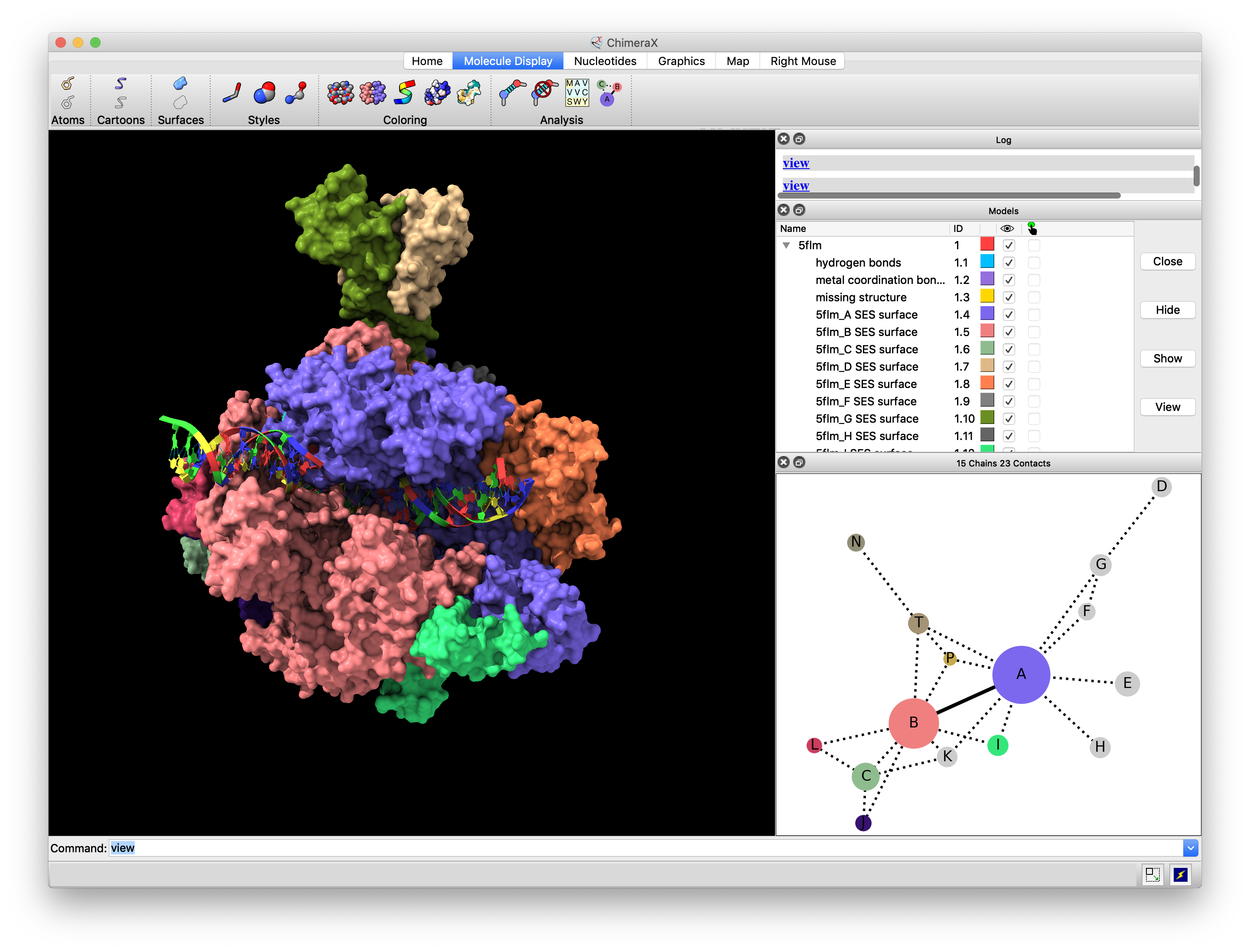
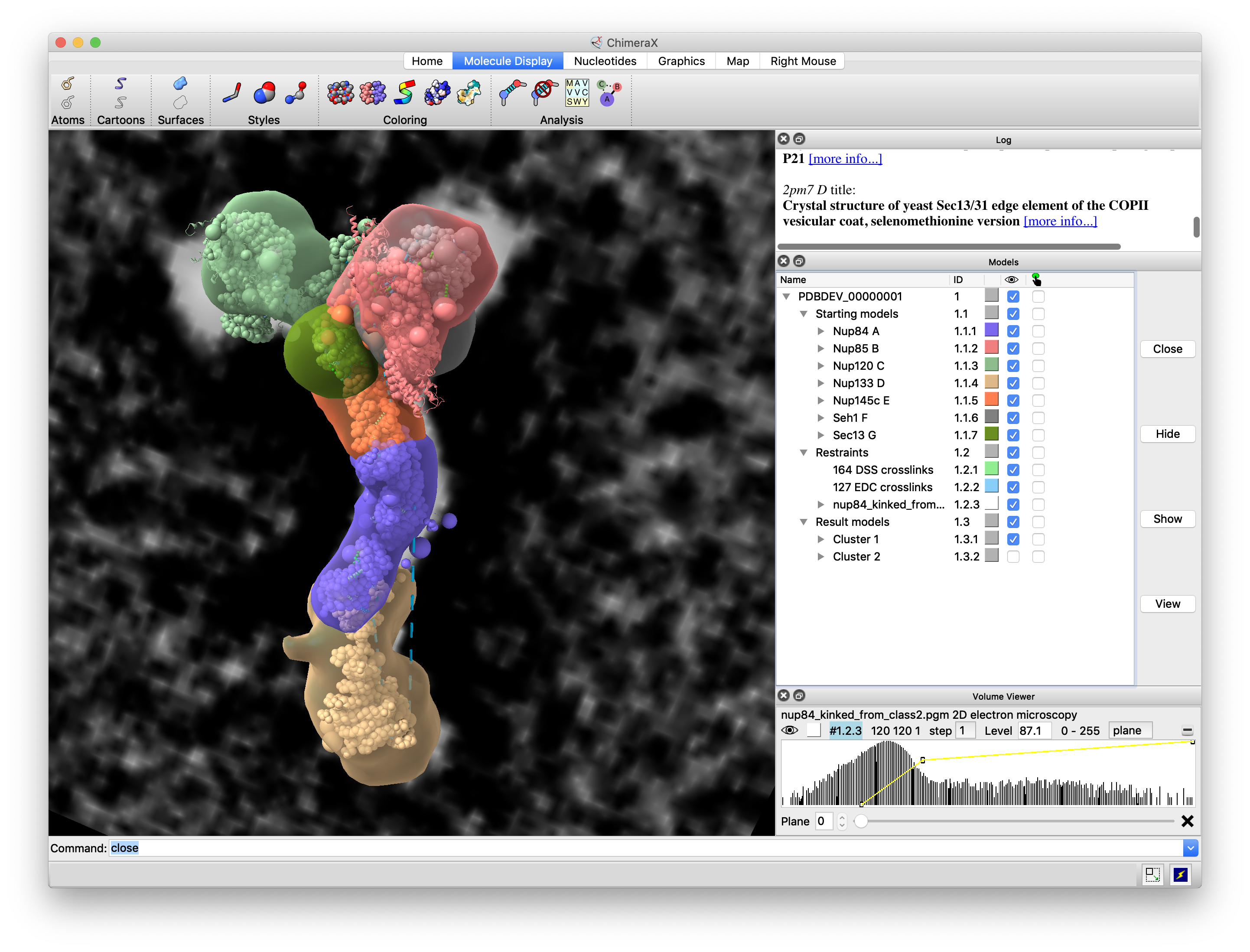
Our Projects
Tool:
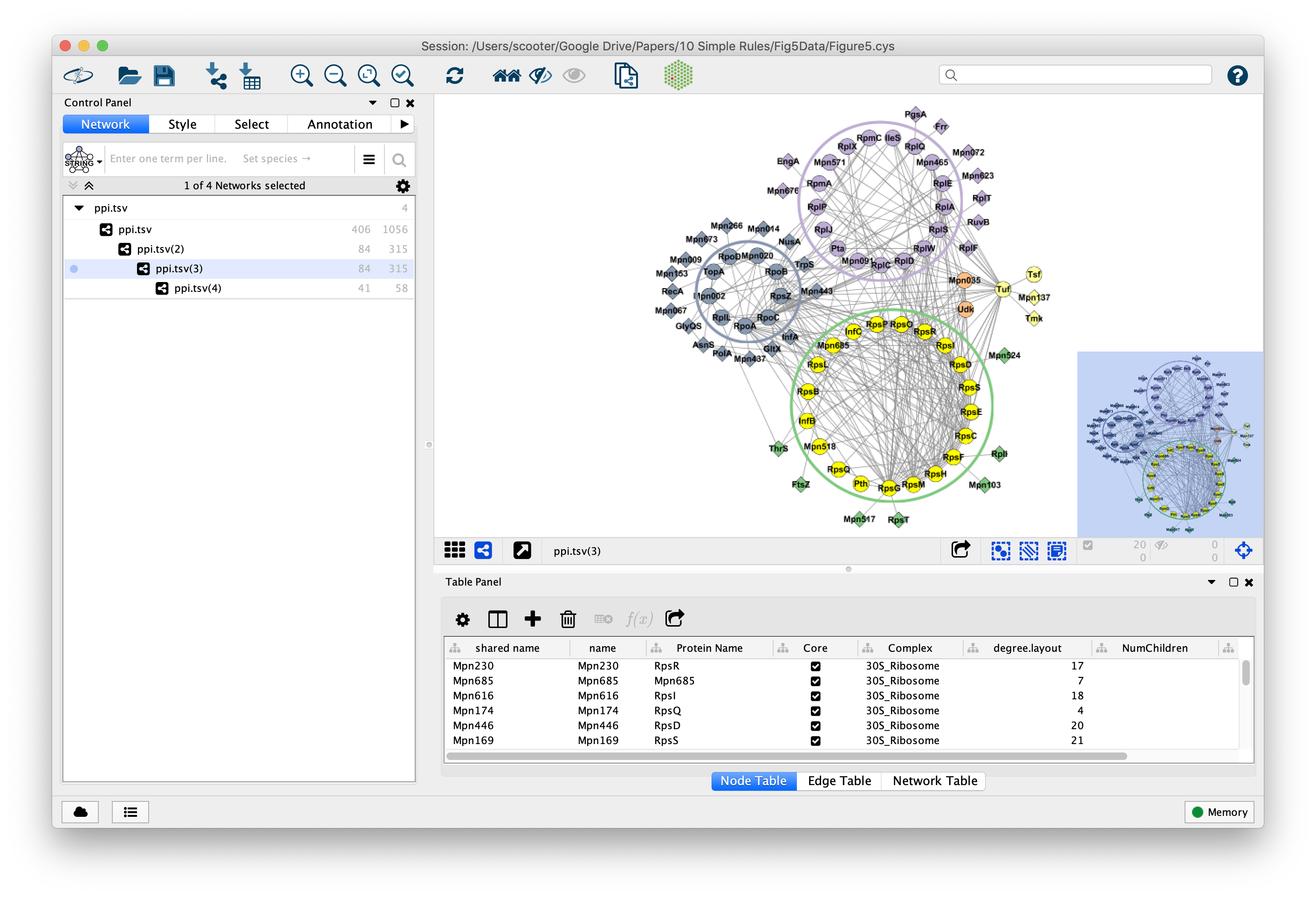
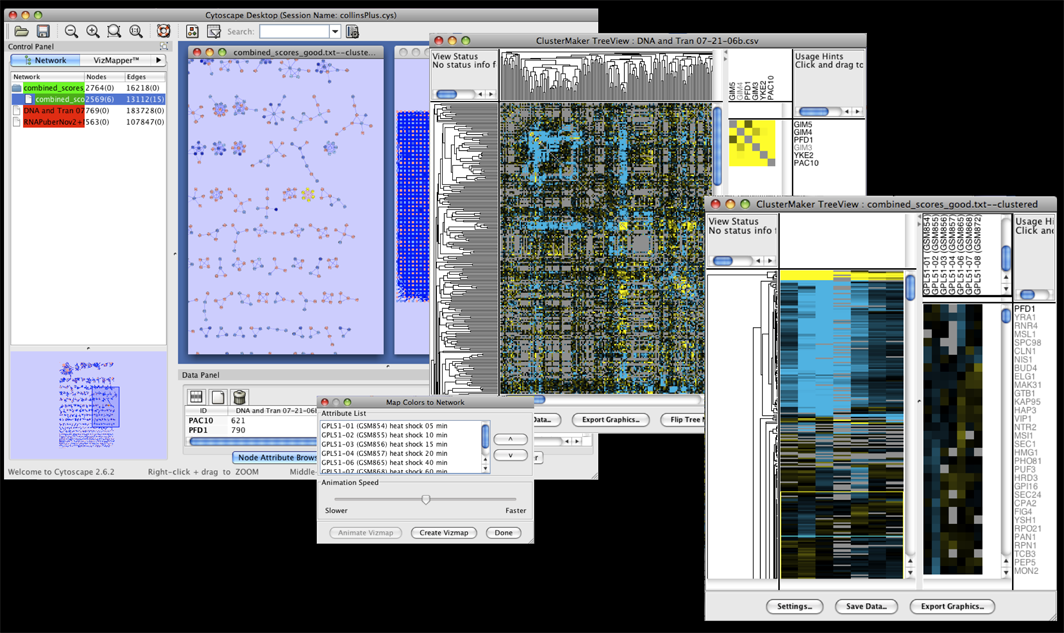
Our Projects
Tool:
Problem : putting single-cell RNA seq results in a broader biological contextChallenge : 100,000 cells, 27,000 genesSolution : aggregate within clusters and do network analysis between clusters
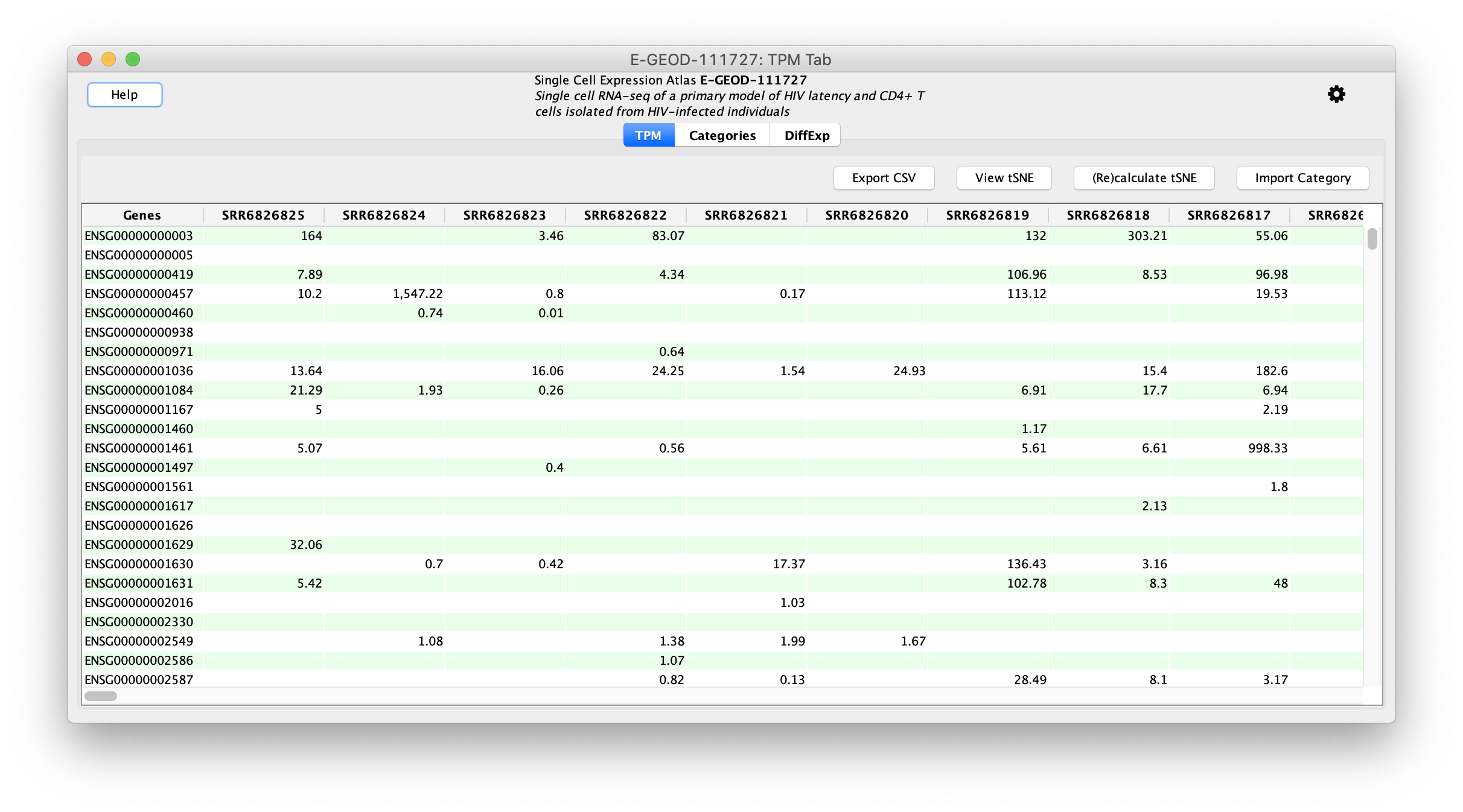
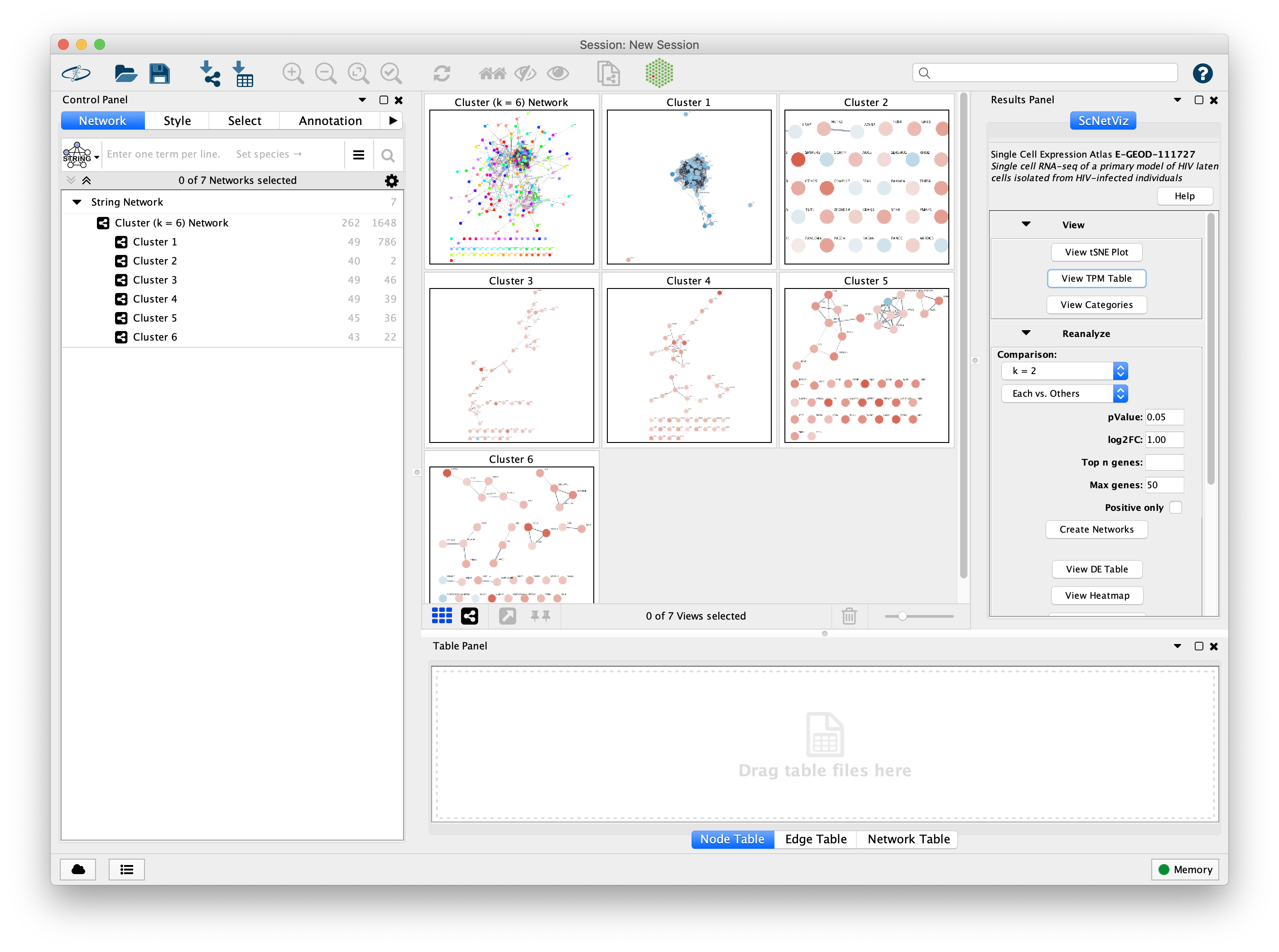
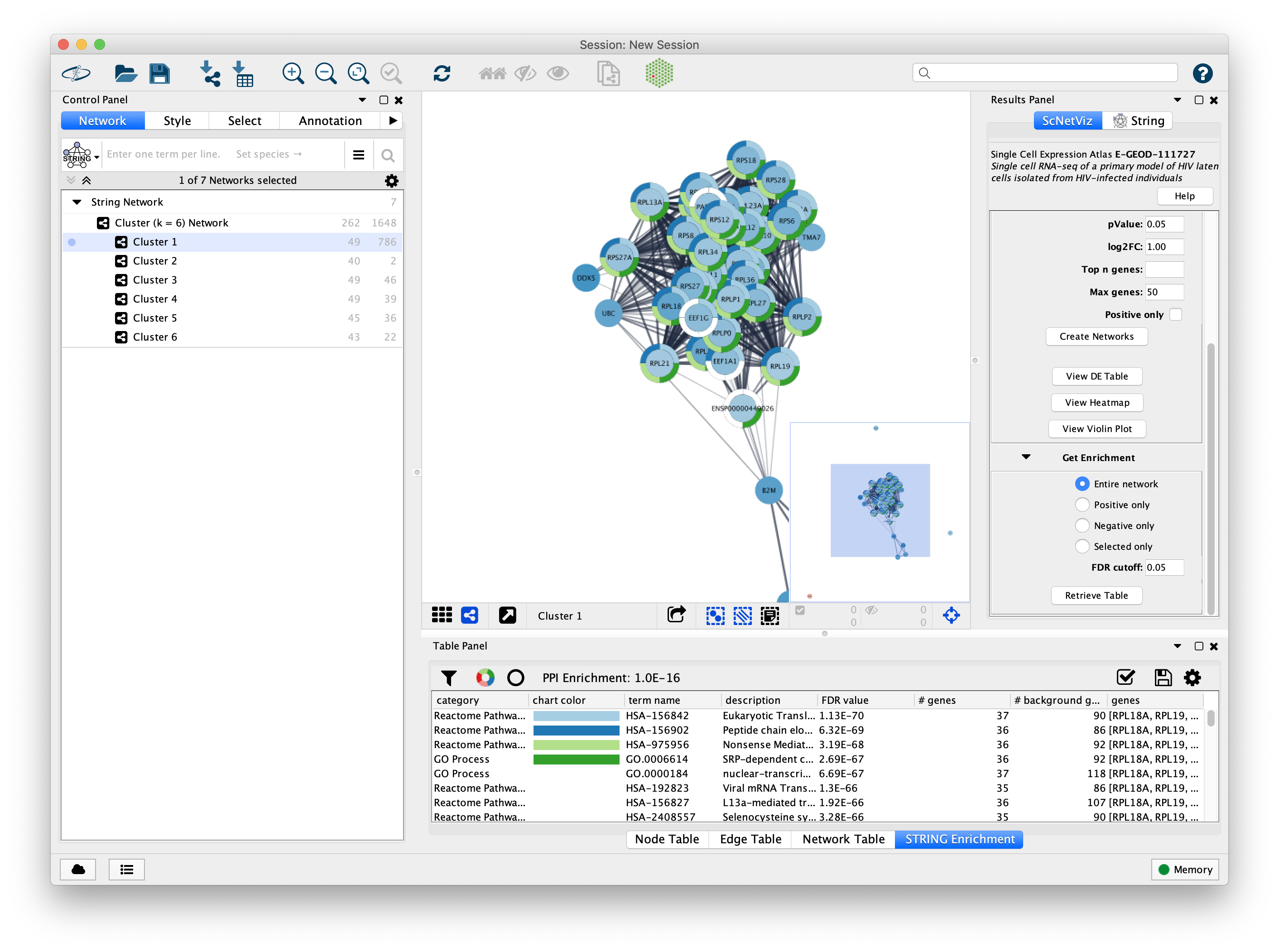
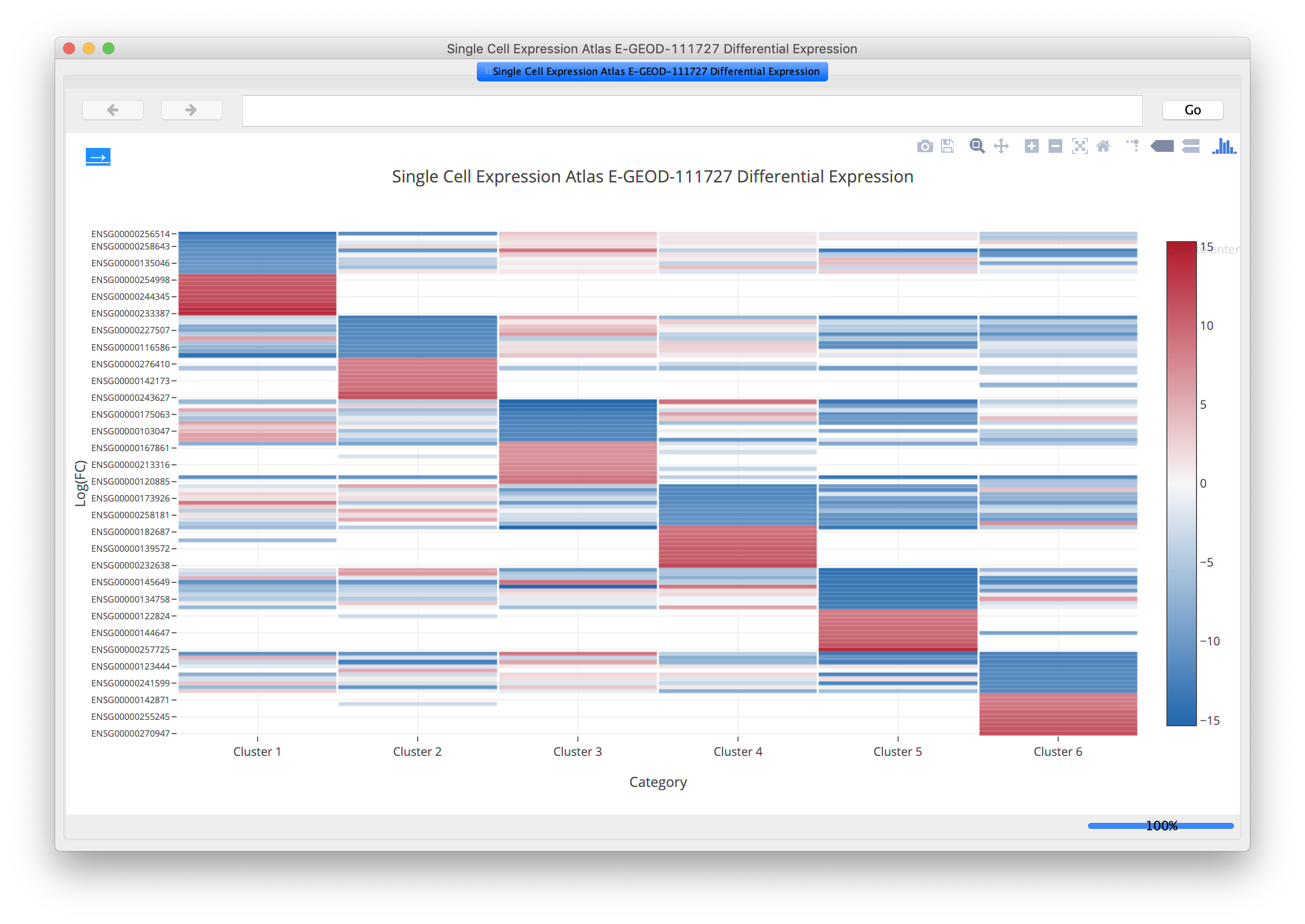
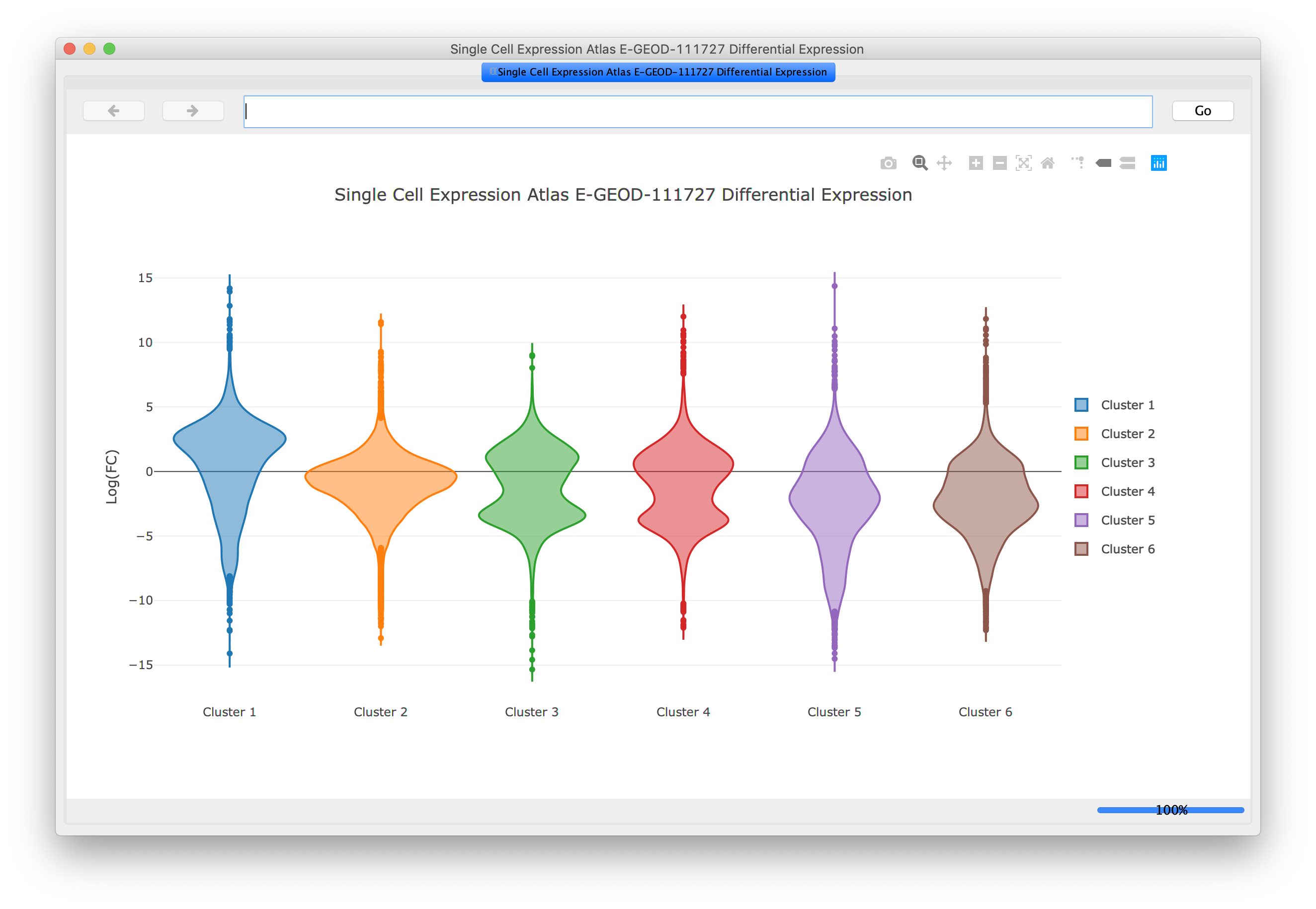
Our Projects
Tool:
Problem : lots of public databases relevant to human diseaseChallenge : they aren't linked and there's no easy way to navigate through themSolution : create a large "knowledge network" that combines multiple databases
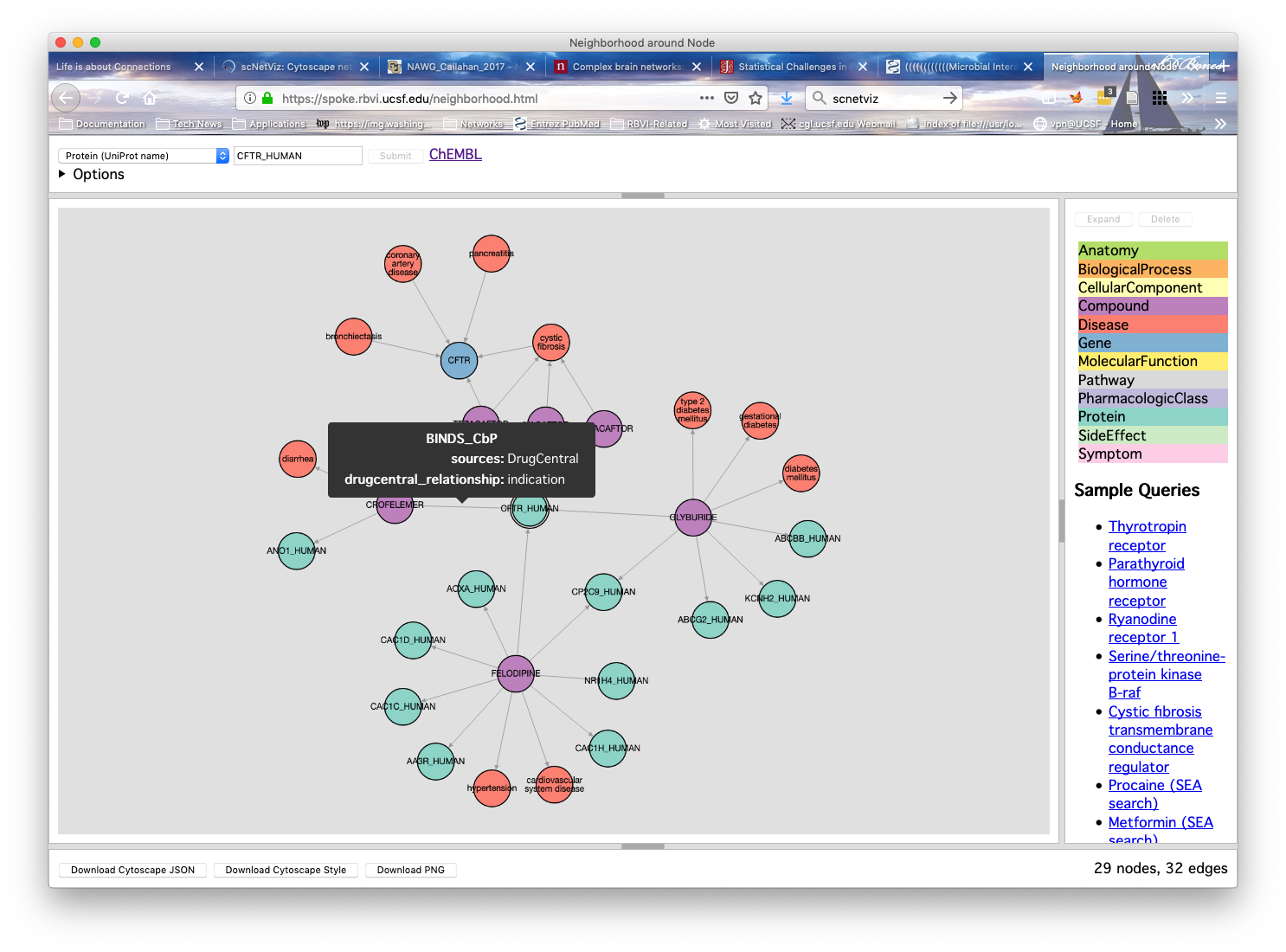
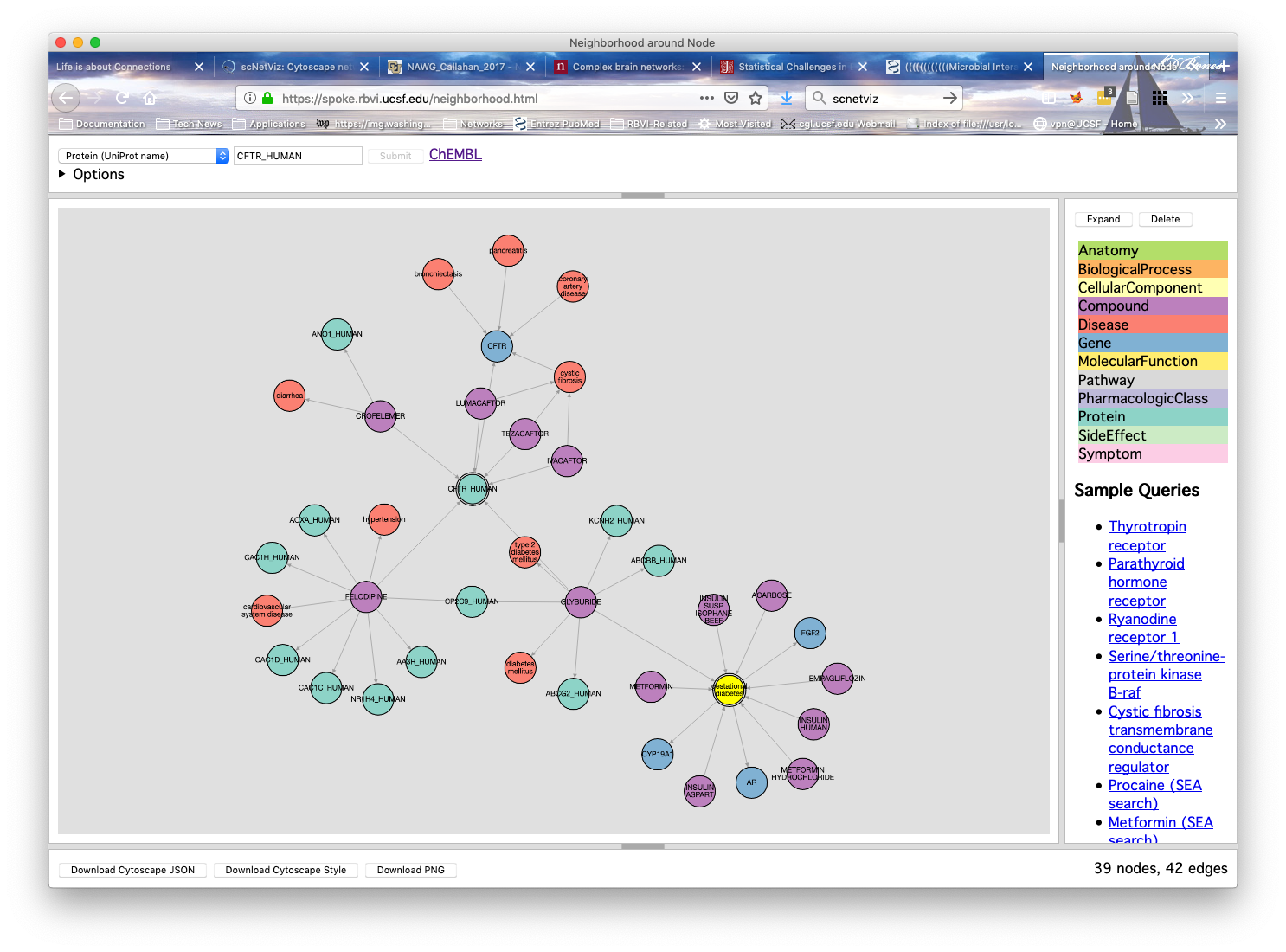
Our Projects
- Anatomy
- BiologicalProcess
- CellularComponent
- Compound
- Disease
- Gene
- MolecularFunction
- Pathway
- PharmacologicClass
- Protein
- SideEffect
- Symptom
- Anatomy→contains→Anatomy
- Anatomy-expresses-Gene
- Anatomy→isa→Anatomy
- Anatomy→partof→Anatomy
- Compound-affects-(mutant)Gene
- Compound-binds-Protein
- Compound-causes-SideEffect
- Compound-contraindicates-Disease
- Compound-resembles-Compound
- Compound-treats-Disease
- Disease-associates-Gene
- Disease→contains→Disease
- Disease→isa→Disease
- Disease-localizes-Anatomy
- Disease-presents-Symptom
- Disease-resembles-Disease
- Gene-covaries-Gene
- Gene-participates-BiologicalProcess
- Gene-participates-CellularComponent
- Gene-participates-MolecularFunction
- Gene-participates-Pathway
- PharmacologicClass-includes-Compound
- Protein-interacts-Protein
- Protein-translatedfrom-Gene
Our Projects
Tool:
- Differential networks
- Dynamic networks
Challenge
Understanding biological connections is critical to:
- understanding biological processes
- understanding how species adapt
- understanding evolution
- understanding (and combating) disease
Challenge
Understanding biological connections will take:
- new computational algorithms and approaches to deal with massive scale
- new approaches to visualization to support in the interpretation of data
- new collaborations between biology, medical science, computer science, engineering, and mathematics
Challenge
- Swansea University has:
- Amazing biology
- Exciting medical sciences
- Talented mathemeticians and computer scientists
- Swansea University doesn't have:
- Computational Biology program to bring all of this together
Challenge
- a computational biology colloquium series, in collaboration with the biomaths colloquia?
- more cross-campus events to bring researchers together?
- an open-door policy to encourage collaboration?


Connections