ChimeraX Recipes
Show CAVER tunnels
Here is how to show CAVER tunnels in biomolecular structures in ChimeraX. We’ll look at example data caver_1bl8.zip that comes with CAVER 3.0 for potassium channel PDB 1bl8. CAVER produces a PDB file for each tunnel in the directory
results/data/clusters_timeless
with names
tun_cl_001_1.pdb
tun_cl_002_1.pdb
...
tun_cl_034_1.pdb
tun_cl_035_1.pdb
We can open all those in ChimeraX, then change the atoms to sphere style.
open 1bl8
open ~/Downloads/caver_1bl8/results/data/clusters_timeless/tun*.pdb
style #2-36 sphere
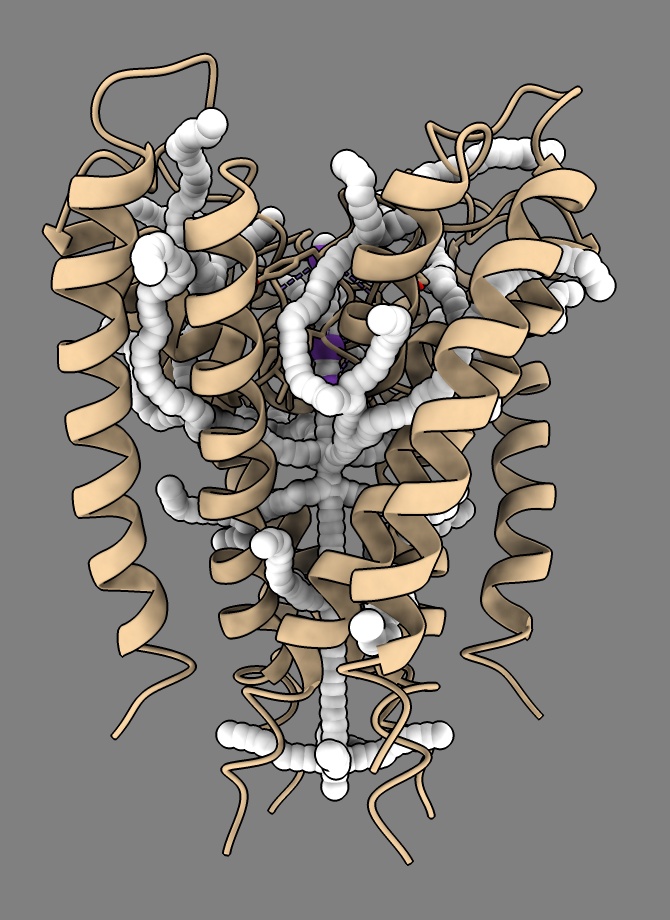
The tunnel radius for each atom is given as the bfactor. ChimeraX does not have a command to set the radius to the b-factor value right now, but we can do it with some Python. First select all the tunnels
select #2-36
Then open this Python file set_radius_to_bfactor.py to set the radii of all the selected atoms to the b-factor
open ~/Downloads/set_radius_to_bfactor.py
Alternatively you can type the following Python code into the ChimeraX Python shell, menu Tools / General / Shell
from chimerax.atomic import selected_atoms
for a in selected_atoms(session):
a.radius = a.bfactor
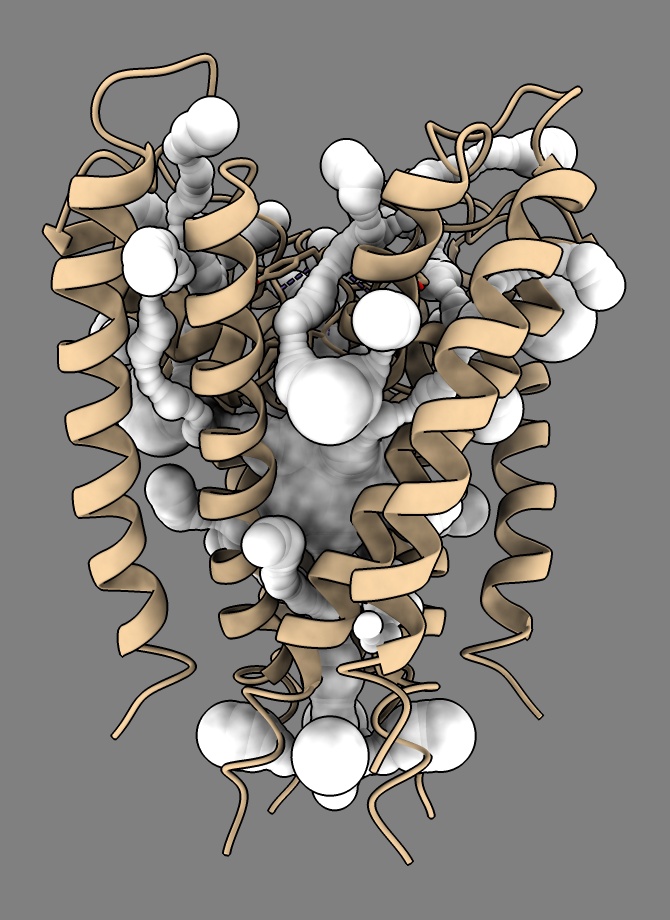
You can color each tunnel differently with
rainbow #2-36 structures
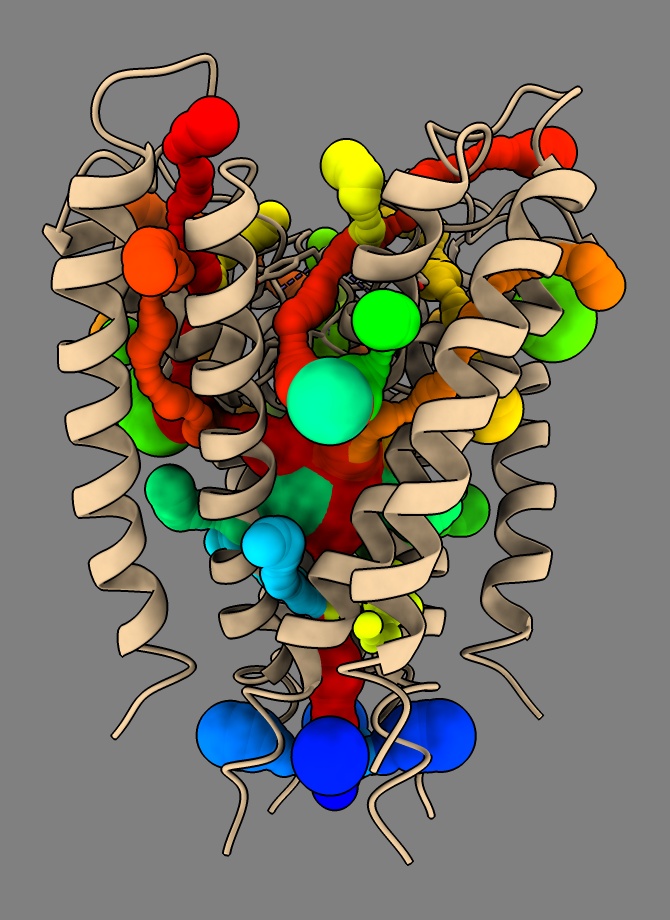
You can flip through them showing each individually with
mseries slider #2-36
The tunnels also look nicer as surfaces instead of atoms
surface #2-36
hide #2-36 atoms
If you hover your mouse over the central channel tunnel you see it is model #23. I can show just that one and color it by electrostatic potential of the protein as in Elaine’s MOLE tutorial (https://www.rbvi.ucsf.edu/chimerax/data/mole-channel/mole-channel.html)
hide #!2-36 models
show #23 model
coulombic protein surfaces #23 offset -1
Also to see the side-chains that come close to the central tunnel I can use the clashes command
clashes #23 restrict #1 overlapCutoff -1 reveal true
That reshows the tunnel atoms so hide them again
hide #23 atoms
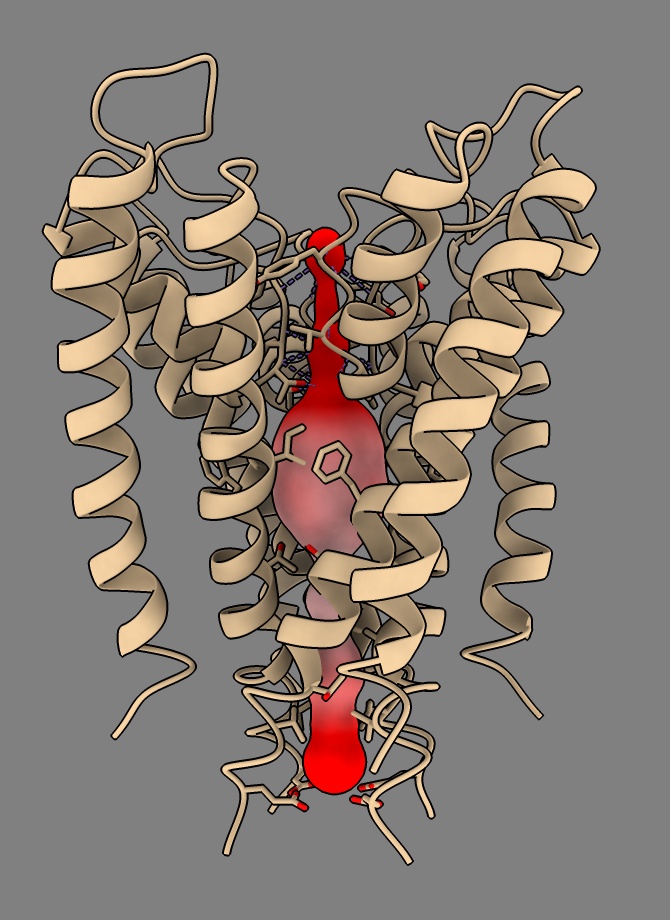
Elaine’s ChimeraX MOLE tutorial also shows electrostatic surfaces on the protein, and she gave a journal-club review of tunnel programs.
Tom Goddard, October 3, 2023